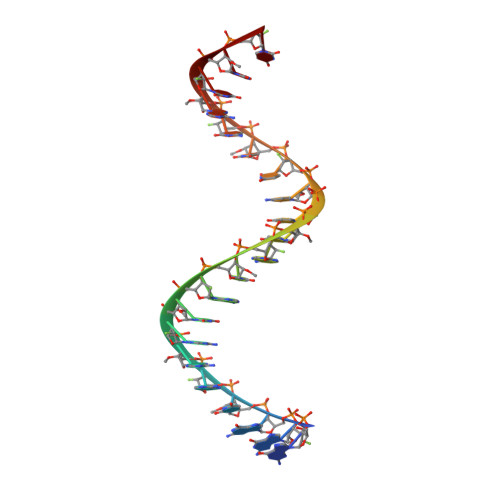
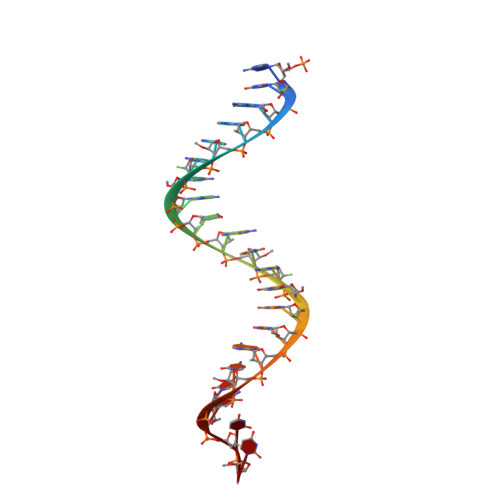
Solution-state structure of a fully alternately 2'-F/2'-OMe modified 42-nt dimeric siRNA construct.
Podbevsek, P., Allerson, C.R., Bhat, B., Plavec, J.(2010) Nucleic Acids Res 38: 7298-7307
- PubMed: 20624819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkq621
- Primary Citation Related Structures:
2KWG - PubMed Abstract:
A high-resolution solution structure of a stable 42-nt RNA dimeric construct has been derived based on a high number of NMR observables including nuclear overhauser effects (NOEs), J-coupling constants and residual dipolar couplings (RDCs), which were all obtained with isotopically unlabeled molecules. Two 21-nt siRNA that efficiently hybridize consist of ribose units that were alternately substituted by 2'-fluoro or 2'-methoxy groups. Structure calculations utilized a set of H-F RDC values for all 21 2'-fluoro modified nucleotides under conditions of weak alignment achieved by Pf1 phages. A completely 2'-F/2'-OMe modified dimeric RNA construct adopts an antiparallel double-helical structure consisting of 19 Watson-Crick base pairs with additional 3' UU overhangs and a 5' phosphate group on the antisense strand. NMR data suggest that the stability of individual base pairs is not uniform throughout the construct. While most of the double helical segment exhibits well dispersed imino resonances, the last three base pairs either display uncharacteristic chemical shifts of imino protons or absence of imino resonances even at lower temperatures. Accessibility of imino protons to solvent exchange suggests a difference in stability of duplex ends, which might be of importance for incorporation of the guide siRNA strand into a RISC.
- Slovenian NMR Center, National Institute of Chemistry, Hajdrihova 19, SI-1001 Ljubljana, Slovenia.
Organizational Affiliation: