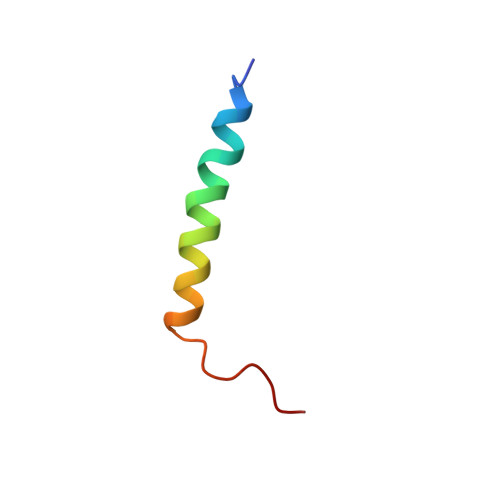
Solution structure and membrane binding of the toxin fst of the par addiction module
Gobl, C., Kosol, S., Stockner, T., Ruckert, H.M., Zangger, K.(2010) Biochemistry 49: 6567-6575
- PubMed: 20677831 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi1005128
- Primary Citation Related Structures:
2KV5 - PubMed Abstract:
The par toxin-antitoxin system is required for the stable inheritance of the plasmid pAD1 in its native host Enterococcus faecalis. It codes for the toxin Fst and a small antisense RNA which inhibits translation of toxin mRNA, and it is the only known antisense regulated toxin-antitoxin system in Gram-positive bacteria. This study presents the structure of the par toxin Fst, the first atomic resolution structure of a component of an antisense regulated toxin-antitoxin system. The mode of membrane binding was determined by relaxation enhancements in a paramagnetic environment and molecular dynamics simulation. Fst forms a membrane-binding alpha-helix in the N-terminal part and contains an intrinsically disordered region near the C-terminus. It binds in a transmembrane orientation with the C-terminus likely pointing toward the cytosol. Membrane-bound, alpha-helical peptides are frequently found in higher organisms as components of the innate immune system. Despite similarities to these antimicrobial peptides, Fst shows neither hemolytic nor antimicrobial activity when applied externally to a series of bacteria, fungal cells, and erythrocytes. Moreover, its charge distribution, orientation in the membrane, and structure distinguish it from antimicrobial peptides.
- Institute of Chemistry/Organic and Bioorganic Chemistry, University of Graz, Heinrichstrasse 28, A-8010 Graz, Austria.
Organizational Affiliation: