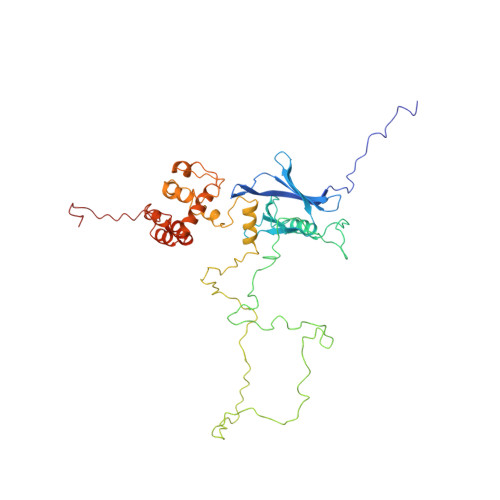
Structure of Proteasome Ubiquitin Receptor hRpn13 and Its Activation by the Scaffolding Protein hRpn2.
Chen, X., Lee, B.H., Finley, D., Walters, K.J.(2010) Mol Cell 38: 404-415
- PubMed: 20471946 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2010.04.019
- Primary Citation Related Structures:
2KQZ, 2KR0 - PubMed Abstract:
Rpn13 is a subunit of the proteasome that serves as a receptor for both ubiquitin and Uch37, one of the proteasome's three deubiquitinating enzymes. We have determined the structure of full-length human Rpn13 (hRpn13). Unexpectedly, Rpn13's ubiquitin- and Uch37-binding domains pack against each other when it is not incorporated into the proteasome. This intramolecular interaction reduces hRpn13's affinity for ubiquitin. We find that hRpn13 binding to the proteasome scaffolding protein hRpn2/S1 abrogates its interdomain interactions, thus activating hRpn13 for ubiquitin binding. hRpn13's Uch37-binding domain, a previously unknown fold, contains nine alpha helices. We have mapped its Uch37-binding surface to a region rich in charged amino acids. Altogether, our results provide mechanistic insights into hRpn13's functional activities with Uch37 and ubiquitin and suggest that its role as a ubiquitin receptor is finely tuned for proteasome targeting.
- Department of Biochemistry, University of Minnesota, Minneapolis, MN 55455, USA.
Organizational Affiliation: