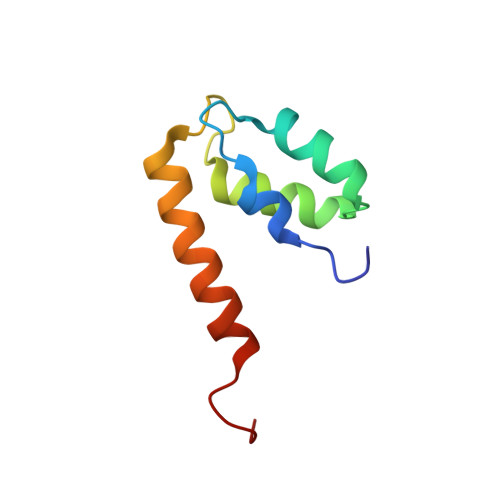
Structure of the EF-hand domain of polycystin-2 suggests a mechanism for Ca2+-dependent regulation of polycystin-2 channel activity.
Petri, E.T., Celic, A., Kennedy, S.D., Ehrlich, B.E., Boggon, T.J., Hodsdon, M.E.(2010) Proc Natl Acad Sci U S A 107: 9176-9181
- PubMed: 20439752 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0912295107
- Primary Citation Related Structures:
2KQ6 - PubMed Abstract:
The C-terminal cytoplasmic tail of polycystin-2 (PC2/TRPP2), a Ca(2+)-permeable channel, is frequently mutated or truncated in autosomal dominant polycystic kidney disease. We have previously shown that this tail consists of three functional regions: an EF-hand domain (PC2-EF, 720-797), a flexible linker (798-827), and an oligomeric coiled coil domain (828-895). We found that PC2-EF binds Ca(2+) at a single site and undergoes Ca(2+)-dependent conformational changes, suggesting it is an essential element of Ca(2+)-sensitive regulation of PC2 activity. Here we describe the NMR structure and dynamics of Ca(2+)-bound PC2-EF. Human PC2-EF contains a divergent non-Ca(2+)-binding helix-loop-helix (HLH) motif packed against a canonical Ca(2+)-binding EF-hand motif. This HLH motif may have evolved from a canonical EF-hand found in invertebrate PC2 homologs. Temperature-dependent steady-state NOE experiments and NMR R(1) and R(2) relaxation rates correlate with increased molecular motion in the EF-hand, possibly due to exchange between apo and Ca(2+)-bound states, consistent with a role for PC2-EF as a Ca(2+)-sensitive regulator. Structure-based sequence conservation analysis reveals a conserved hydrophobic surface in the same region, which may mediate Ca(2+)-dependent protein interactions. We propose that Ca(2+)-sensing by PC2-EF is responsible for the cooperative nature of PC2 channel activation and inhibition. Based on our results, we present a mechanism of regulation of the Ca(2+) dependence of PC2 channel activity by PC2-EF.
- Department of Pharmacology, Yale University School of Medicine, 333 Cedar Street, New Haven, CT 06520, USA.
Organizational Affiliation: