An NMR-based structural model of the PthA repeat region reveals a TPR fold that would account for protein-protein and protein-DNA interactions
Neves, J.L., Sforca, M.L., Murakami, M.T., Benedetti, C.E., Zeri, A.C.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Avirulence protein | 51 | Xanthomonas citri | Mutation(s): 0 | 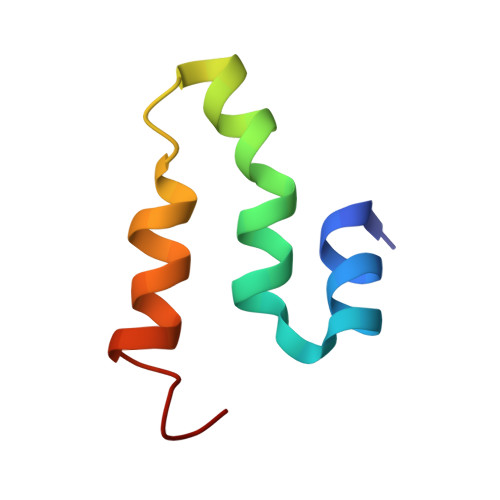 | |