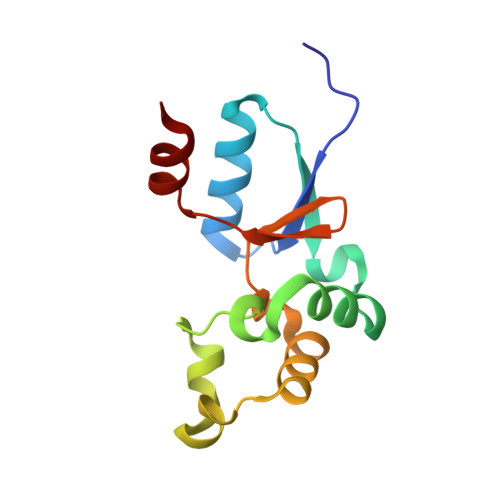
Solution structure of an arsenate reductase-related protein, YffB, from Brucella melitensis, the etiological agent responsible for brucellosis.
Buchko, G.W., Hewitt, S.N., Napuli, A.J., Van Voorhis, W.C., Myler, P.J.(2011) Acta Crystallogr Sect F Struct Biol Cryst Commun 67: 1129-1136
- PubMed: 21904062 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309111006336
- Primary Citation Related Structures:
2KOK - PubMed Abstract:
Brucella melitensis is the etiological agent responsible for brucellosis. Present in the B. melitensis genome is a 116-residue protein related to arsenate reductases (Bm-YffB; BR0369). Arsenate reductases (ArsC) convert arsenate ion (H(2)AsO(4)(-)), a compound that is toxic to bacteria, to arsenite ion (AsO(2)(-)), a product that may be efficiently exported out of the cell. Consequently, Bm-YffB is a potential drug target because if arsenate reduction is the protein's major biological function then disabling the cell's ability to reduce arsenate would make these cells more sensitive to the deleterious effects of arsenate. Size-exclusion chromatography and NMR spectroscopy indicate that Bm-YffB is a monomer in solution. The solution structure of Bm-YffB (PDB entry 2kok) shows that the protein consists of two domains: a four-stranded mixed β-sheet flanked by two α-helices on one side and an α-helical bundle. The α/β domain is characteristic of the fold of thioredoxin-like proteins and the overall structure is generally similar to those of known arsenate reductases despite the marginal sequence similarity. Chemical shift perturbation studies with (15)N-labeled Bm-YffB show that the protein binds reduced glutathione at a site adjacent to a region similar to the HX(3)CX(3)R catalytic sequence motif that is important for arsenic detoxification activity in the classical arsenate-reductase family of proteins. The latter observation supports the hypothesis that the ArsC-YffB family of proteins may function as glutathione-dependent thiol reductases. However, comparison of the structure of Bm-YffB with the structures of proteins from the classical ArsC family suggest that the mechanism and possibly the function of Bm-YffB and other related proteins (ArsC-YffB) may differ from those of the ArsC family of proteins.
- Seattle Structural Genomics Center for Infectious Disease, http://www.ssgcid.org, USA. garry.buchko@pnnl.gov
Organizational Affiliation: