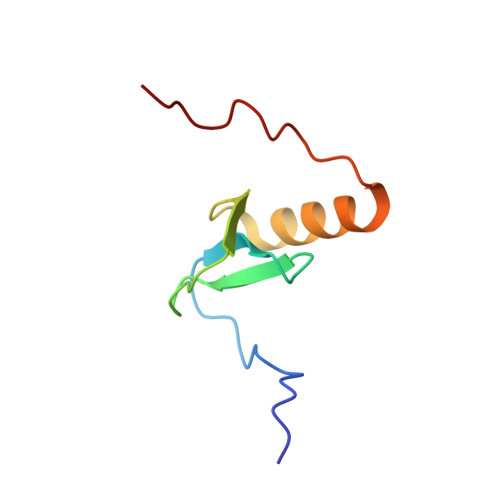
Solution structure of Rv2377c-founding member of the MbtH-like protein family.
Buchko, G.W., Kim, C.Y., Terwilliger, T.C., Myler, P.J.(2010) Tuberculosis (Edinb) 90: 245-251
- PubMed: 20434955 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.tube.2010.04.002
- Primary Citation Related Structures:
2KHR - PubMed Abstract:
The Mycobacterium tuberculosis protein Rv2377c (71 residues, MW=8.4kDa) has been characterized using nuclear magnetic resonance (NMR) and circular dichroism (CD) spectroscopy. Rv2377c was the first identified member of the MbtH-like family of proteins. MbtH-like proteins have been implicated in siderophore biosynthesis, however, their precise biochemical function remain unknown. Size exclusion chromatography and NMR spectroscopy show that Rv2377c is a monomer in solution. Circular dichroism spectroscopy indicates that Rv2377c unfolds upon heating and will reversibly fold into its native conformation upon cooling. Using NMR-based methods the solution structure of Rv2377c was determined and some of the dynamic properties of the protein studied. The protein contains a three-strand, anti-parallel beta-sheet (beta3:beta1:beta2) nestled against one C-terminal alpha-helix (S44-N55). Weak or absent amide cross peaks in the (1)H-(15)N HSQC spectrum for many of the beta1 and beta2 residues suggest intermediate motion on the ms to mus time scale at the beta1:beta2 interface. Amide cross peaks in the (1)H-(15)N HSQC spectrum are absent for all but one residue at the C-terminus (W56-D71), a region that includes a highly conserved sequence WXDXR, suggesting this region is intrinsically disordered. The latter observation differs with the crystal structure of another MbtH-like protein, PA2412 from Pseudomonas aeruginosa, where a second ordered alpha-helix was observed at the extreme C-terminus.
- Pacific Northwest National Laboratory, Richland, WA 99352, USA. garry.buchko@pnl.gov
Organizational Affiliation: