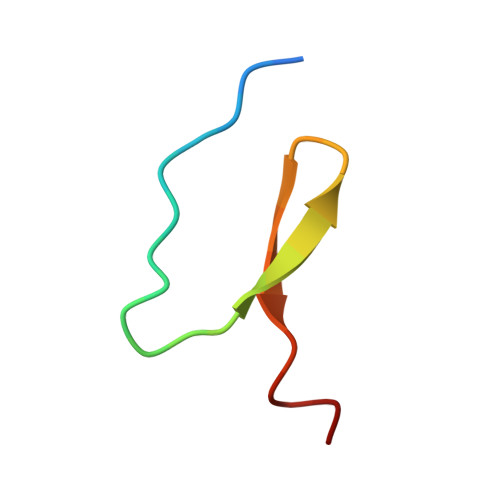
NMR Structure of a Monomeric Intermediate on the Evolutionarily Optimized Assembly Pathway of a Small Trimerization Domain
Habazettl, J., Reiner, A., Kiefhaber, T.(2009) J Mol Biology 389: 103-114
- PubMed: 19361528 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.03.073
- Primary Citation Related Structures:
2KBL - PubMed Abstract:
Efficient formation of specific intermolecular interactions is essential for self-assembly of biological structures. The foldon domain is an evolutionarily optimized trimerization module required for assembly of the large, trimeric structural protein fibritin from phage T4. Monomers consisting of the 27 amino acids comprising a single foldon domain subunit spontaneously form a natively folded trimer. During assembly of the foldon domain, a monomeric intermediate is formed on the submillisecond time scale, which provides the basis for two consecutive very fast association reactions. Mutation of an intermolecular salt bridge leads to a monomeric protein that resembles the kinetic intermediate in its spectroscopic properties. NMR spectroscopy revealed essentially native topology of the monomeric intermediate with defined hydrogen bonds and side-chain interactions but largely reduced stability compared to the native trimer. This structural preorganization leads to an asymmetric charge distribution on the surface that can direct rapid subunit recognition. The low stability of the intermediate allows a large free-energy gain upon trimerization, which serves as driving force for rapid assembly. These results indicate different free-energy landscapes for folding of small oligomeric proteins compared to monomeric proteins, which typically avoid the transient population of intermediates.
- Biozentrum der Universität Basel, Switzerland.
Organizational Affiliation: