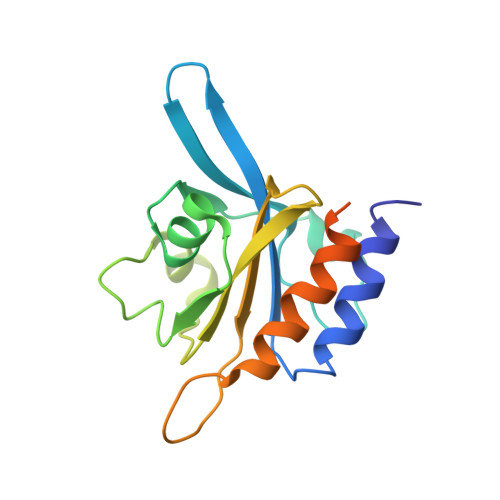
Solution Structure of the cGMP Binding GAF Domain from Phosphodiesterase 5: Insights into Nucleotide Selectivity, Dimerization, and cGMP-Dependent Conformational Change.
Heikaus, C.C., Stout, J.R., Sekharan, M.R., Eakin, C.M., Rajagopal, P., Brzovic, P.S., Beavo, J.A., Klevit, R.E.(2008) J Biological Chem 283: 22749-22759
- PubMed: 18534985 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M801577200
- Primary Citation Related Structures:
2K31 - PubMed Abstract:
Phosphodiesterase 5 (PDE5) controls intracellular levels of cGMP through its regulation of cGMP hydrolysis. Hydrolytic activity of the C-terminal catalytic domain is increased by cGMP binding to the N-terminal GAF A domain. We present the NMR solution structure of the cGMP-bound PDE5A GAF A domain. The cGMP orientation in the buried binding pocket was defined through 37 intermolecular nuclear Overhauser effects. Comparison with GAF domains from PDE2A and adenylyl cyclase cyaB2 reveals a conserved overall domain fold of a six-stranded beta-sheet and four alpha-helices that form a well defined cGMP binding pocket. However, the nucleotide coordination is distinct with a series of altered binding contacts. The structure suggests that nucleotide binding specificity is provided by Asp-196, which is positioned to form two hydrogen bonds to the guanine ring of cGMP. An alanine mutation of Asp-196 disrupts cGMP binding and increases cAMP affinity in constructs containing only GAF A causing an altered cAMP-bound structural conformation. NMR studies on the tandem GAF domains reveal a flexible GAF A domain in the absence of cGMP, and indicate a large conformational change upon ligand binding. Furthermore, we identify a region of approximately 20 residues directly N-terminal of GAF A as critical for tight dimerization of the tandem GAF domains. The features of the PDE5 regulatory domain revealed here provide an initial structural basis for future investigations of the regulatory mechanism of PDE5 and the design of GAF-specific regulators of PDE5 function.
- Department of Biochemistry, University of Washington, Seattle, Washington 98195, USA.
Organizational Affiliation: