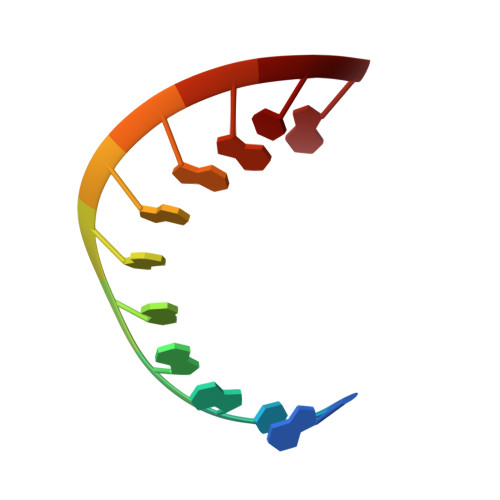
Bulged adenosine influence on the RNA duplex conformation in solution.
Popenda, L., Adamiak, R.W., Gdaniec, Z.(2008) Biochemistry 47: 5059-5067
- PubMed: 18399645
- DOI: https://doi.org/10.1021/bi7024904
- Primary Citation Related Structures:
2JXQ, 2JXS - PubMed Abstract:
The RNA single bulge motif is an unpaired residue within a strand of several complementary base pairs. To gain insight into structural changes induced by the presence of the adenosine bulge on RNA duplex, the solution structures of RNA duplex containing a single adenine bulge (5'-GCAGAAGAGCG-3'/5'-CGCUCUCUGC-3') and a reference duplex with all Watson-Crick base pairs (5'-GCAGAGAGCG-3'/5'-CGCUCUCUGC-3') have been determined by NMR spectroscopy. The reference duplex structure is a regular right-handed helix with all of the attributes of an A-type helix. In the bulged duplex, single adenine bulge stacks into the helix, and the bulge region forms a well-defined structure. Both structures were analyzed by the use of calculated helical parameters. Distortions induced by the accommodation of unpaired residue into the helical structure propagate over the entire structure and are manifested as the reduced base pairs inclination and x-displacement. Intrahelical position of bulged adenine A5 is stabilized by efficient stacking with 5'-neighboring residues G4.
- Institute of Bioorganic Chemistry, Polish Academy of Science, Noskowskiego 12/14, 61-704 Poznań, Poland.
Organizational Affiliation: