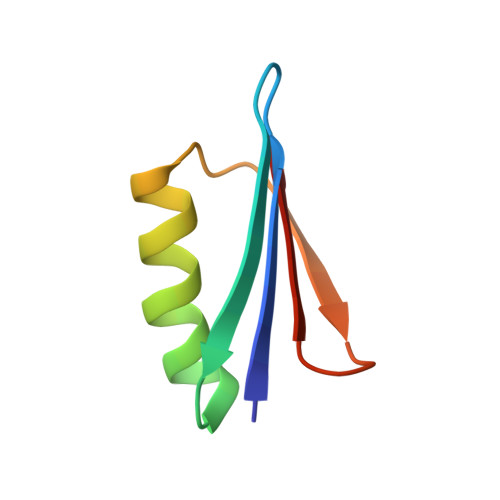
NMR structures of two designed proteins with high sequence identity but different fold and function
He, Y., Chen, Y., Alexander, P., Bryan, P.N., Orban, J.(2008) Proc Natl Acad Sci U S A 105: 14412-14417
- PubMed: 18796611 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0805857105
- Primary Citation Related Structures:
2JWS, 2JWU - PubMed Abstract:
How protein sequence codes for 3D structure remains a fundamental question in biology. One approach to understanding the folding code is to design a pair of proteins with maximal sequence identity but retaining different folds. Therefore, the nonidentities must be responsible for determining which fold topology prevails and constitute a fold-specific folding code. We recently designed two proteins, G(A)88 and G(B)88, with 88% sequence identity but different folds and functions [Alexander et al. (2007) Proc Natl Acad Sci USA 104:11963-11968]. Here, we describe the detailed 3D structures of these proteins determined in solution by NMR spectroscopy. Despite a large number of mutations taking the sequence identity level from 16 to 88%, G(A)88 and G(B)88 maintain their distinct wild-type 3-alpha and alpha/beta folds, respectively. To our knowledge, the 3D-structure determination of two monomeric proteins with such high sequence identity but different fold topology is unprecedented. The geometries of the seven nonidentical residues (of 56 total) provide insights into the structural basis for switching between 3-alpha and alpha/beta conformations. Further mutation of a subset of these nonidentities, guided by the G(A)88 and G(B)88 structures, leads to proteins with even higher levels of sequence identity (95%) and different folds. Thus, conformational switching to an alternative monomeric fold of comparable stability can be effected with just a handful of mutations in a small protein. This result has implications for understanding not only the folding code but also the evolution of new folds.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, MD 20850, USA.
Organizational Affiliation: