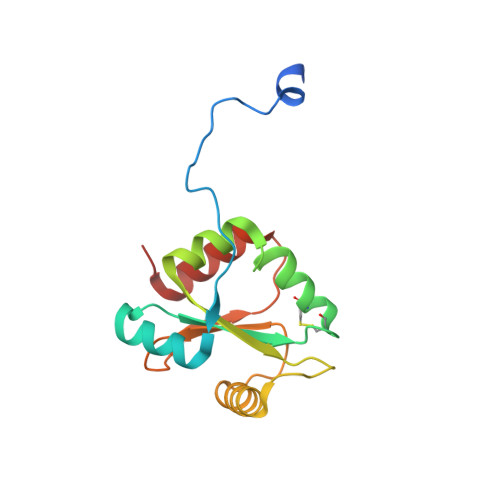
Insight into Disulfide Bond Catalysis in Chlamydia from the Structure and Function of DsbH, a Novel Oxidoreductase.
Mac, T.T., von Hacht, A., Hung, K.C., Dutton, R.J., Boyd, D., Bardwell, J.C., Ulmer, T.S.(2008) J Biological Chem 283: 824-832
- PubMed: 18003611 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M707863200
- Primary Citation Related Structures:
2JU5 - PubMed Abstract:
The Chlamydia family of human pathogens uses outer envelope proteins that are highly cross-linked by disulfide bonds but nevertheless keeps an unusually high number of unpaired cysteines in its secreted proteins. To gain insight into chlamydial disulfide bond catalysis, the structure, function, and substrate interaction of a novel periplasmic oxidoreductase, termed DsbH, were determined. The structure of DsbH, its redox potential of -269 mV, and its functional properties are similar to thioredoxin and the C-terminal domain of DsbD, i.e. characteristic of a disulfide reductase. As compared with these proteins, the two central residues of the DsbH catalytic motif (CMWC) shield the catalytic disulfide bond and are selectively perturbed by a peptide ligand. This shows that these oxidoreductase family characteristic residues are not only important in determining the redox potential of the catalytic disulfide bond but also in influencing substrate interactions. For DsbH, three functional roles are conceivable; that is, reducing intermolecular disulfides between proteins and small molecules, keeping a specific subset of exported proteins reduced, or maintaining the periplasm of Chlamydia in a generally reducing state.
- Department of Biochemistry and Molecular Biology and Zilkha Neurogenetic Institute, Keck School of Medicine, University of Southern California, Los Angeles, California 90033, USA.
Organizational Affiliation: