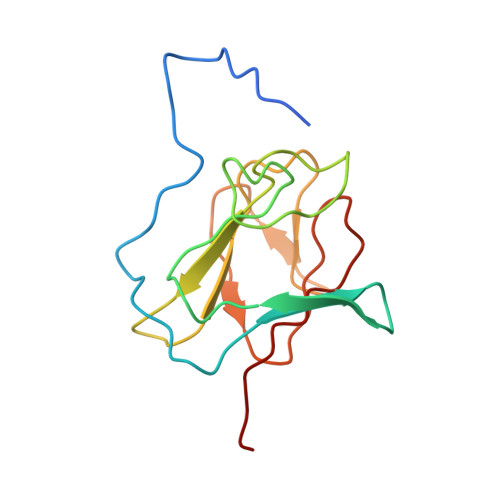
The NMR structure of the NIPP1 FHA domain.
Kumeta, H., Ogura, K., Adachi, S., Fujioka, Y., Tanuma, N., Tanuma, K., Kikuchi, K., Inagaki, F.(2008) J Biomol NMR 40: 219-224
- PubMed: 18253837 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-008-9222-x
- Primary Citation Related Structures:
2JPE - Graduate School of Pharmaceutical Sciences, Hokkaido University, Sapporo, Hokkaido, 060-0812, Japan.
Organizational Affiliation: