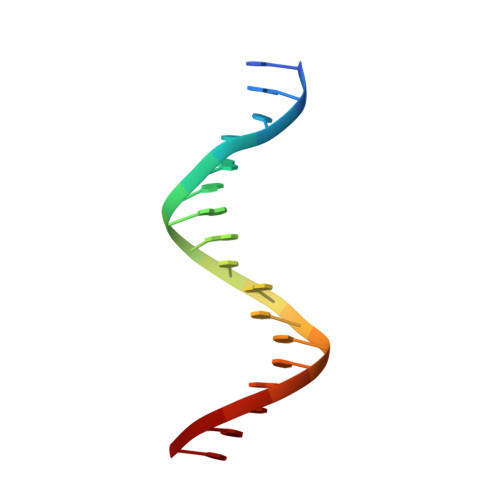
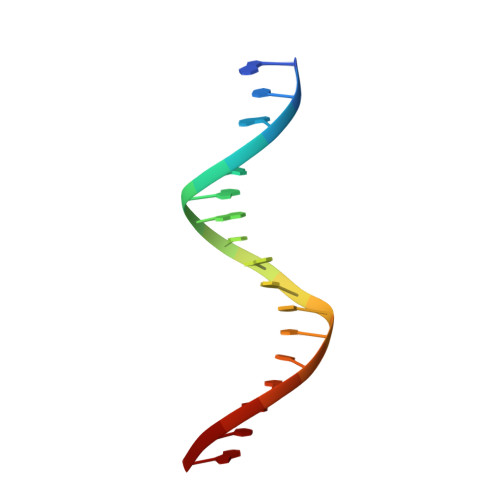
Structure of the wilms tumor suppressor protein zinc finger domain bound to DNA
Stoll, R., Lee, B.M., Debler, E.W., Laity, J.H., Wilson, I.A., Dyson, H.J., Wright, P.E.(2007) J Mol Biology 372: 1227-1245
- PubMed: 17716689 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.07.017
- Primary Citation Related Structures:
2JP9, 2JPA, 2PRT - PubMed Abstract:
The zinc finger domain of the Wilms tumor suppressor protein (WT1) contains four canonical Cys(2)His(2) zinc fingers. WT1 binds preferentially to DNA sequences that are closely related to the EGR-1 consensus site. We report the structure determination by both X-ray crystallography and NMR spectroscopy of the WT1 zinc finger domain in complex with DNA. The X-ray structure was determined for the complex with a cognate 14 base-pair oligonucleotide, and composite X-ray/NMR structures were determined for complexes with both the 14 base-pair and an extended 17 base-pair DNA. This combined approach allowed unambiguous determination of the position of the first zinc finger, which is influenced by lattice contacts in the crystal structure. The crystal structure shows the second, third and fourth zinc finger domains inserted deep into the major groove of the DNA where they make base-specific interactions. The DNA duplex is distorted in the vicinity of the first zinc finger, with a cytidine twisted and tilted out of the base stack to pack against finger 1 and the tip of finger 2. By contrast, the composite X-ray/NMR structures show that finger 1 continues to follow the major groove in the solution complexes. However, the orientation of the helix is non-canonical, and the fingertip and the N terminus of the helix project out of the major groove; as a consequence, the zinc finger side-chains that are commonly involved in base recognition make no contact with the DNA. We conclude that finger 1 helps to anchor WT1 to the DNA by amplifying the binding affinity although it does not contribute significantly to binding specificity. The structures provide molecular level insights into the potential consequences of mutations in zinc fingers 2 and 3 that are associated with Denys-Drash syndrome and nephritic syndrome. The mutations are of two types, and either destabilize the zinc finger structure or replace key base contact residues.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: