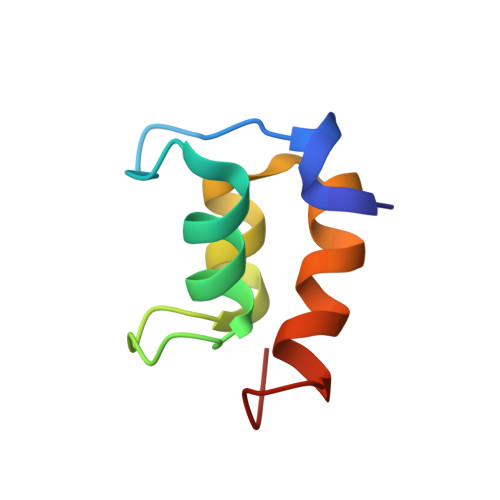
Cold-adaptation in Sea-water-borne Signal Proteins: Sequence and NMR Structure of the Pheromone En-6 from the Antarctic Ciliate Euplotes nobilii
Pedrini, B., Placzek, W.J., Koculi, E., Alimenti, C., LaTerza, A., Luporini, P., Wuthrich, K.(2007) J Mol Biology 372: 277-286
- PubMed: 17663000
- DOI: https://doi.org/10.1016/j.jmb.2007.06.046
- Primary Citation of Related Structures:
2JMS - PubMed Abstract:
Ciliates of Euplotes species constitutively secrete pleiotropic protein pheromones, which are capable to function as prototypic autocrine growth factors as well as paracrine inducers of mating processes. This paper reports the amino acid sequence and the NMR structure of the pheromone En-6 isolated from the antarctic species Euplotes nobilii. The 63-residue En-6 polypeptide chain forms three alpha-helices in positions 18-25, 36-40 and 46-56, which are arranged in an up-down-up three-helix bundle forming the edges of a distorted trigonal pyramid. The base of the pyramid is covered by the N-terminal heptadecapeptide segment, which includes a 3(10)-turn of residues 3-6. This topology is covalently anchored by four long-range disulfide bonds. Comparison with the smaller pheromones of E. raikovi, a closely related species living in temperate waters, shows that the two-pheromone families have the same three-helix bundle architecture. It then appears that cold-adaptation of the En proteins is primarily related to increased lengths of the chain-terminal peptide segments and the surface-exposed loops connecting the regular secondary structures, and to the presence of solvent-exposed clusters of negatively charged side-chains.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: