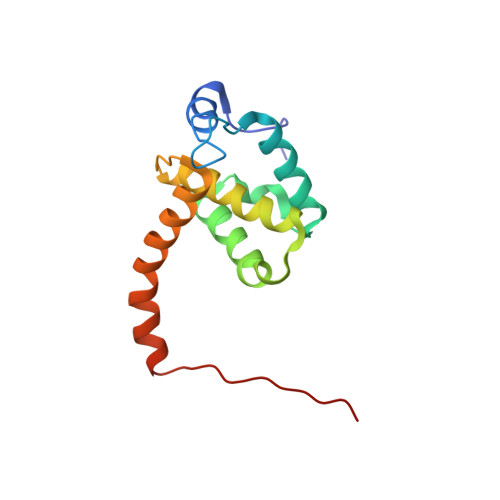
Point mutations in the HIV-1 matrix protein turn off the myristyl switch.
Saad, J.S., Loeliger, E., Luncsford, P., Liriano, M., Tai, J., Kim, A., Miller, J., Joshi, A., Freed, E.O., Summers, M.F.(2007) J Mol Biology 366: 574-585
- PubMed: 17188710 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.11.068
- Primary Citation Related Structures:
2JMG, 2NV3 - PubMed Abstract:
During the late phase of human immunodeficiency virus type-1 (HIV-1) replication, newly synthesized retroviral Gag proteins are targeted to lipid raft regions of specific cellular membranes, where they assemble and bud to form new virus particles. Gag binds preferentially to the plasma membrane (PM) of most hematopoietic cell types, a process mediated by interactions between the cellular PM marker phosphatidylinositol-(4,5)-bisphosphate (PI(4,5)P(2)) and Gag's N-terminally myristoylated matrix (MA) domain. We recently demonstrated that PI(4,5)P(2) binds to a conserved cleft on MA and promotes myristate exposure, suggesting a role as both a direct membrane anchor and myristyl switch trigger. Here we show that PI(4,5)P(2) is also capable of binding to MA proteins containing point mutations that inhibit membrane binding in vitro, and in vivo, including V7R, L8A and L8I. However, these mutants do not exhibit PI(4,5)P(2) or concentration-dependent myristate exposure. NMR studies of V7R and L8A MA reveal minor structural changes that appear to be responsible for stabilizing the myristate-sequestered (myr(s)) species and inhibiting exposure. Unexpectedly, the myristyl group of a revertant mutant with normal PM targeting properties (V7R,L21K) is also tightly sequestered and insensitive to PI(4,5)P(2) binding. This mutant binds PI(4,5)P(2) with twofold higher affinity compared with the native protein, suggesting a potential compensatory mechanism for membrane binding.
- Howard Hughes Medical Institute and Department of Chemistry and Biochemistry, University of Maryland Baltimore County, 1000 Hilltop Circle, Baltimore, MD 21250, USA.
Organizational Affiliation: