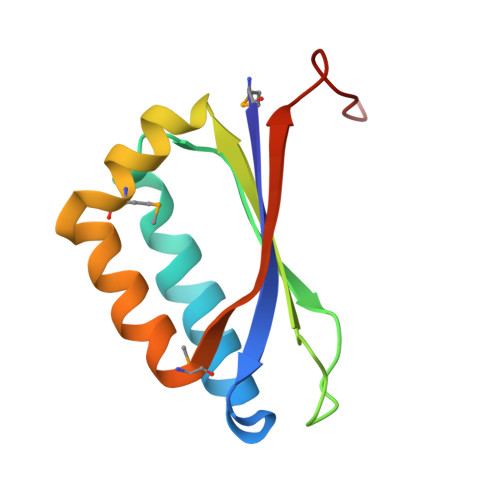
Structural Insight of the Role of the Hahella Chejuensis Hapk Protein in Prodigiosin Biosynthesis.
Cho, H.J., Kim, K., Kim, M.H., Kang, B.S.(2008) Proteins 70: 257
- PubMed: 17729271
- DOI: https://doi.org/10.1002/prot.21582
- Primary Citation of Related Structures:
2JDJ - School of Life Science and Biotechnology, Kyungpook National University, Daegu 702-701, Korea.
Organizational Affiliation: