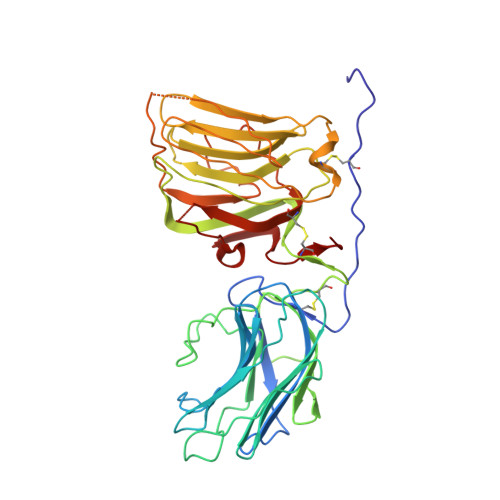
Crystal Structure and Cell Surface Anchorage Sites of Laminin {Alpha}1Lg4-5.
Harrison, D., Hussain, S.A., Combs, A.C., Ervasti, J.M., Yurchenco, P.D., Hohenester, E.(2007) J Biological Chem 282: 11573
- PubMed: 17307732 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M610657200
- Primary Citation Related Structures:
2JD4 - PubMed Abstract:
The laminin G-like (LG) domains of laminin-111, a glycoprotein widely expressed during embryogenesis, provide cell anchoring and receptor binding sites that are involved in basement membrane assembly and cell signaling. We now report the crystal structure of the laminin alpha1LG4-5 domains and provide a mutational analysis of heparin, alpha-dystroglycan, and galactosylsulfatide binding. The two domains of alpha1LG4-5 are arranged in a V-shaped fashion similar to that observed with laminin alpha2 LG4-5 but with a substantially different interdomain angle. Recombinant alpha1LG4-5 binding to heparin, alpha-dystroglycan, and sulfatides was dependent upon both shared and unique contributions from basic residues distributed in several clusters on the surface of LG4. For heparin, the greatest contribution was detected from two clusters, 2719RKR and 2791KRK. Binding to alpha-dystroglycan was particularly dependent on basic residues within 2719RKR, 2831RAR, and 2858KDR. Binding to galactosylsulfatide was most affected by mutations in 2831RAR and 2766KGRTK but not in 2719RKR. The combined analysis of structure and activities reveal differences in LG domain interactions that should enable dissection of biological roles of different laminin ligands.
- Department of Pathology and Laboratory Medicine, Robert Wood Johnson Medical School, Piscataway, New Jersey 08854, USA.
Organizational Affiliation: