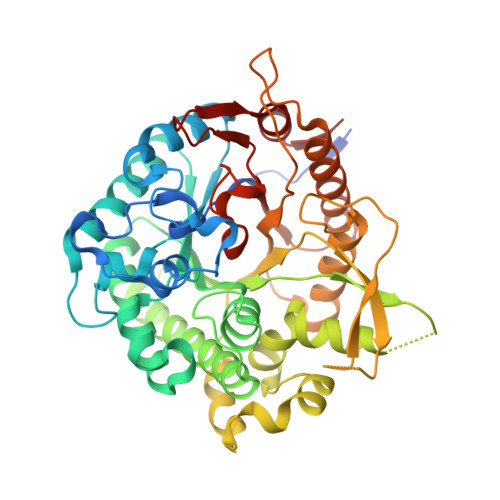
Structural Basis for Cyclophellitol Inhibition of a Beta-Glucosidase.
Gloster, T.M., Madsen, R., Davies, G.J.(2007) Org Biomol Chem 5: 444
- PubMed: 17252125 Search on PubMed
- DOI: https://doi.org/10.1039/b616590g
- Primary Citation Related Structures:
2JAL - PubMed Abstract:
The structural basis for beta-glucosidase inhibition by cyclophellitol is demonstrated using X-ray crystallography, enzyme kinetics and mass spectrometry.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York, UK YO10 5YW.
Organizational Affiliation: