Factor Xa Inhibitors: S1 Binding Interactions of a Series of N-{(3S)-1-[(1S)-1-Methyl-2-Morpholin-4-Yl-2-Oxoethyl]-2-Oxopyrrolidin-3-Yl}Sulfonamides.
Chan, C., Borthwick, A.D., Brown, D., Burns-Kurtis, C.L., Campbell, M., Chaudry, L., Chung, C.W., Convery, M.A., Hamblin, J.N., Johnstone, L., Kelly, H.A., Kleanthous, S., Patikis, A., Patel, C., Pateman, A.J., Senger, S., Shah, G.P., Toomey, J.R., Watson, N.S., Weston, H.E., Whitworth, C., Young, R.J., Zhou, P.(2007) J Med Chem 50: 1546
- PubMed: 17338508 Search on PubMed
- DOI: https://doi.org/10.1021/jm060870c
- Primary Citation Related Structures:
2J94, 2J95, 4Y76, 4Y79 - PubMed Abstract:
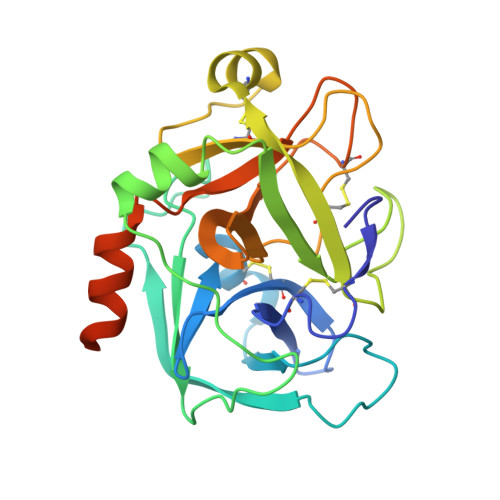
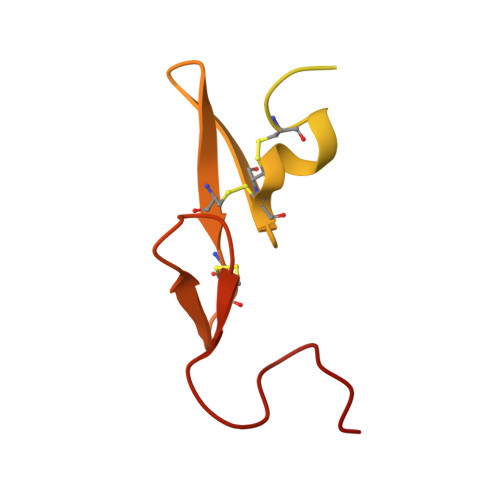
Factor Xa inhibitory activities for a series of N-{(3S)-1-[(1S)-1-methyl-2-morpholin-4-yl-2-oxoethyl]-2-oxopyrrolidin-3-yl}sulfonamides with different P1 groups are described. These data provide insight into binding interactions within the S1 primary specificity pocket; rationales are presented for the derived SAR on the basis of electronic interactions through crystal structures of fXa-ligand complexes and molecular modeling studies. A good correlation between in vitro anticoagulant activities with lipophilicity and the extent of human serum albumin binding is observed within this series of potent fXa inhibitors. Pharmacokinetic profiles in rat and dog, together with selectivity over other trypsin-like serine proteases, identified 1f as a candidate for further evaluation.
- CVU UK Medicinal Chemistry Department, GlaxoSmithKline Research and Development, Medicines Research Centre, Gunnels Wood Road, Stevenage, Hertfordshire SG1 2NY, United Kingdom. chuen.8chan@gsk.com
Organizational Affiliation: