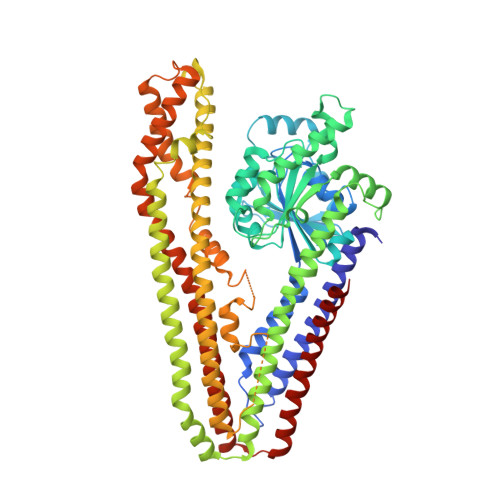
A Bacterial Dynamin-Like Protein
Low, H.H., Lowe, J.(2006) Nature 444: 766
- PubMed: 17122778
- DOI: https://doi.org/10.1038/nature05312
- Primary Citation of Related Structures:
2J68, 2J69 - PubMed Abstract:
Dynamins form a superfamily of large mechano-chemical GTPases that includes the classical dynamins and dynamin-like proteins (DLPs). They are found throughout the Eukarya, functioning in core cellular processes such as endocytosis and organelle division. Many bacteria are predicted by sequence to possess large GTPases with the same multidomain architecture that is found in DLPs. Mechanistic dissection of dynamin family members has been impeded by a lack of high-resolution structural data currently restricted to the GTPase and pleckstrin homology domains, and the dynamin-related human guanylate-binding protein. Here we present the crystal structure of a cyanobacterial DLP in both nucleotide-free and GDP-associated conformation. The bacterial DLP shows dynamin-like qualities, such as helical self-assembly and tubulation of a lipid bilayer. In vivo, it localizes to the membrane in a manner reminiscent of FZL, a chloroplast-specific dynamin-related protein with which it shares sequence similarity. Our results provide structural and mechanistic insight that may be relevant across the dynamin superfamily. Concurrently, we show compelling similarity between a cyanobacterial and chloroplast DLP that, given the endosymbiotic ancestry of chloroplasts, questions the evolutionary origins of dynamins.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: