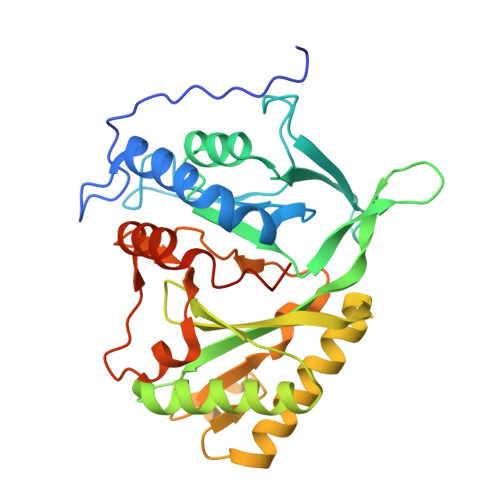
Structure and function of a mycobacterial NHEJ DNA repair polymerase.
Pitcher, R.S., Brissett, N.C., Picher, A.J., Andrade, P., Juarez, R., Thompson, D., Fox, G.C., Blanco, L., Doherty, A.J.(2007) J Mol Biology 366: 391-405
- PubMed: 17174332 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.10.046
- Primary Citation Related Structures:
2IRU, 2IRX, 2IRY - PubMed Abstract:
Non homologous end-joining (NHEJ)-mediated repair of DNA double-strand breaks in prokaryotes requires Ku and a specific multidomain DNA ligase (LigD). We present crystal structures of the primase/polymerisation domain (PolDom) of Mycobacterium tuberculosis LigD, alone and complexed with nucleotides. The PolDom structure combines the general fold of the archaeo-eukaryotic primase (AEP) superfamily with additional loops and domains that together form a deep cleft on the surface, likely used for DNA binding. Enzymatic analysis indicates that the PolDom of LigD, even in the absence of accessory domains and Ku proteins, has the potential to recognise DNA end-joining intermediates. Strikingly, one of the main signals for the specific and efficient binding of PolDom to DNA is the presence of a 5'-phosphate group, located at the single/double-stranded junction at both gapped and 3'-protruding DNA molecules. Although structurally unrelated, Pol lambda and Pol mu, the two eukaryotic DNA polymerases involved in NHEJ, are endowed with a similar capacity to bind a 5'-phosphate group. Other properties that are beneficial for NHEJ, such as the ability to generate template distortions and realignments of the primer, displayed by Pol lambda and Pol mu, are shared by the PolDom of bacterial LigD. In addition, PolDom can perform non-mutagenic translesion synthesis on termini containing modified bases. Significantly, ribonucleotide insertion appears to be a recurrent theme associated with NHEJ, maximised in this case by the deployment of a dedicated primase, although its in vivo relevance is unknown.
- Genome Damage and Stability Centre, University of Sussex, Brighton BN1 9RQ, UK.
Organizational Affiliation: