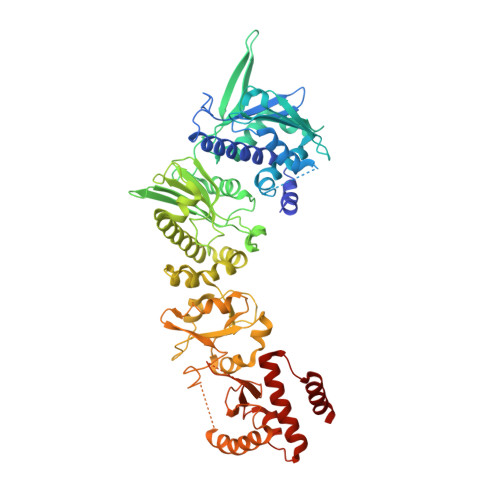
Structural Analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements.
Shiau, A.K., Harris, S.F., Southworth, D.R., Agard, D.A.(2006) Cell 127: 329-340
- PubMed: 17055434 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2006.09.027
- Primary Citation Related Structures:
2GQ0, 2IOP, 2IOQ, 2IOR - PubMed Abstract:
In eukaryotes, the ubiquitous and abundant members of the 90 kilodalton heat-shock protein (hsp90) chaperone family facilitate the folding and conformational changes of a broad array of proteins important in cell signaling, proliferation, and survival. Here we describe the effects of nucleotides on the structure of full-length HtpG, the Escherichia coli hsp90 ortholog. By electron microscopy, the nucleotide-free, AMPPNP bound, and ADP bound states of HtpG adopt completely distinct conformations. Structural characterization of nucleotide-free and ADP bound HtpG was extended to higher resolution by X-ray crystallography. In the absence of nucleotide, HtpG exhibits an "open" conformation in which the three domains of each monomer present hydrophobic elements into the large cleft formed by the dimer. By contrast, ADP binding drives dramatic conformational changes that allow these hydrophobic elements to converge and shield each other from solvent, suggesting a mechanism by which nucleotides could control client protein binding and release.
- Howard Hughes Medical Institute and Department of Biochemistry and Biophysics, University of California, San Francisco, 94158, USA.
Organizational Affiliation: