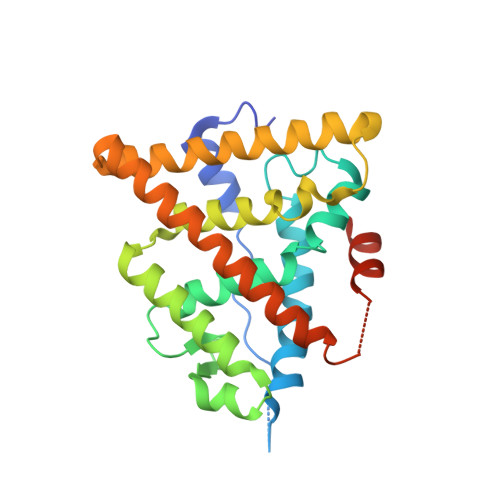
Estrogen receptor ligands. Part 16: 2-Aryl indoles as highly subtype selective ligands for ERalpha
Dykstra, K.D., Guo, L., Birzin, E.T., Chan, W., Yang, Y.T., Hayes, E.C., DaSilva, C.A., Pai, L.-Y., Mosley, R.T., Kraker, B., Fitzgerald, P.M.D., DiNinno, F., Rohrer, S.P., Schaeffer, J.M., Hammond, M.L.(2007) Bioorg Med Chem Lett 17: 2322-2328
- PubMed: 17289385 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2007.01.054
- Primary Citation Related Structures:
2IOG, 2IOK - PubMed Abstract:
A novel class of indole ligands for estrogen receptor alpha have been discovered which exhibit potent affinity and high selectivity. Substitution of the bazedoxifene skeleton to the linker present in the HTS lead 1a provided 22b which was found to be 130-fold alpha-selective and acted as an antagonist of estradiol activity in uterine tissue and MCF-7 cancer cells.
- Department of Medicinal Chemistry, Merck Research Laboratories, PO Box 2000, 800B-109 Rahway, NJ 07065, USA. kevin_dykstra@merck.com
Organizational Affiliation: