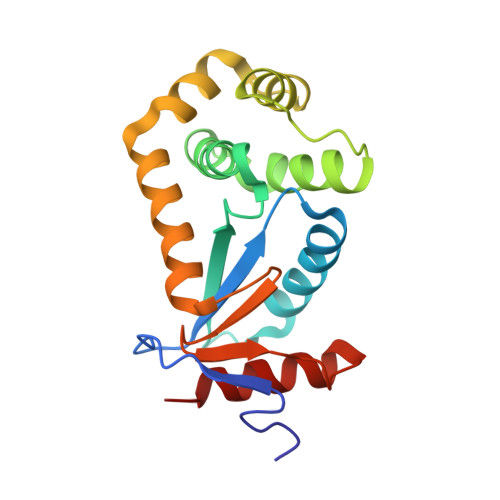
Probing the Flexibility of the DsbA Oxidoreductase from Vibrio cholerae-a (15)N - (1)H Heteronuclear NMR Relaxation Analysis of Oxidized and Reduced Forms of DsbA.
Horne, J., d'Auvergne, E.J., Coles, M., Velkov, T., Chin, Y., Charman, W.N., Prankerd, R., Gooley, P.R., Scanlon, M.J.(2007) J Mol Biology 371: 703-716
- PubMed: 17585933 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.05.067
- Primary Citation Related Structures:
2IJY - PubMed Abstract:
We have determined the structure of the reduced form of the DsbA oxidoreductase from Vibrio cholerae. The reduced structure shows a high level of similarity to the crystal structure of the oxidized form and is typical of this class of enzyme containing a thioredoxin domain with an inserted alpha-helical domain. Proteolytic and thermal stability measurements show that the reduced form of DsbA is considerably more stable than the oxidized form. NMR relaxation data have been collected and analyzed using a model-free approach to probe the dynamics of the reduced and oxidized states of DsbA. Akaike's information criteria have been applied both in the selection of the model-free models and the diffusion tensors that describe the global motions of each redox form. Analysis of the dynamics reveals that the oxidized protein shows increased disorder on the pico- to nanosecond and micro- to millisecond timescale. Many significant changes in dynamics are located either close to the active site or at the insertion points between the domains. In addition, analysis of the diffusion data shows there is a clear difference in the degree of interdomain movement between oxidized and reduced DsbA with the oxidized form being the more rigid. Principal components analysis has been employed to indicate possible concerted movements in the DsbA structure, which suggests that the modeled interdomain motions affect the catalytic cleft of the enzyme. Taken together, these data provide compelling evidence of a role for dynamics in the catalytic cycle of DsbA.
- Department of Medicinal Chemistry, Victorian College of Pharmacy, Monash University, 381 Royal Parade, Parkville, VIC 3052, Australia.
Organizational Affiliation: