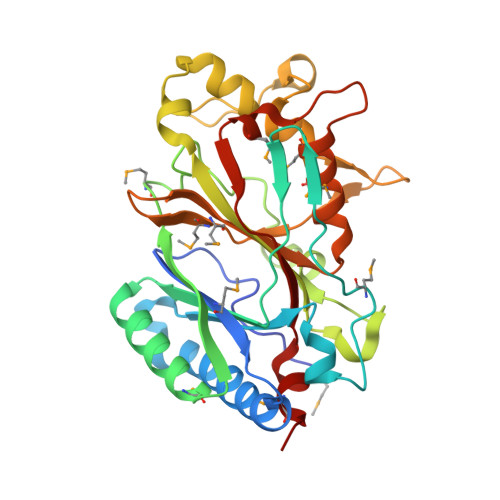
Identification and structural characterization of heme binding in a novel dye-decolorizing peroxidase, TyrA.
Zubieta, C., Joseph, R., Krishna, S.S., McMullan, D., Kapoor, M., Axelrod, H.L., Miller, M.D., Abdubek, P., Acosta, C., Astakhova, T., Carlton, D., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Elias, Y., Elsliger, M.A., Feuerhelm, J., Grzechnik, S.K., Hale, J., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Kumar, A., Marciano, D., Morse, A.T., Murphy, K.D., Nigoghossian, E., Okach, L., Oommachen, S., Reyes, R., Rife, C.L., Schimmel, P., Trout, C.V., van den Bedem, H., Weekes, D., White, A., Xu, Q., Hodgson, K.O., Wooley, J., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2007) Proteins 69: 234-243
- PubMed: 17654547
- DOI: https://doi.org/10.1002/prot.21673
- Primary Citation Related Structures:
2IIZ - PubMed Abstract:
TyrA is a member of the dye-decolorizing peroxidase (DyP) family, a new family of heme-dependent peroxidase recently identified in fungi and bacteria. Here, we report the crystal structure of TyrA in complex with iron protoporphyrin (IX) at 2.3 A. TyrA is a dimer, with each monomer exhibiting a two-domain, alpha/beta ferredoxin-like fold. Both domains contribute to the heme-binding site. Co-crystallization in the presence of an excess of iron protoporphyrin (IX) chloride allowed for the unambiguous location of the active site and the specific residues involved in heme binding. The structure reveals a Fe-His-Asp triad essential for heme positioning, as well as a novel conformation of one of the heme propionate moieties compared to plant peroxidases. Structural comparison to the canonical DyP family member, DyP from Thanatephorus cucumeris (Dec 1), demonstrates conservation of this novel heme conformation, as well as residues important for heme binding. Structural comparisons with representative members from all classes of the plant, bacterial, and fungal peroxidase superfamily demonstrate that TyrA, and by extension the DyP family, adopts a fold different from all other structurally characterized heme peroxidases. We propose that a new superfamily be added to the peroxidase classification scheme to encompass the DyP family of heme peroxidases.
- Stanford Synchrotron Radiation Laboratory, Stanford University, Menlo Park, California, USA.
Organizational Affiliation: