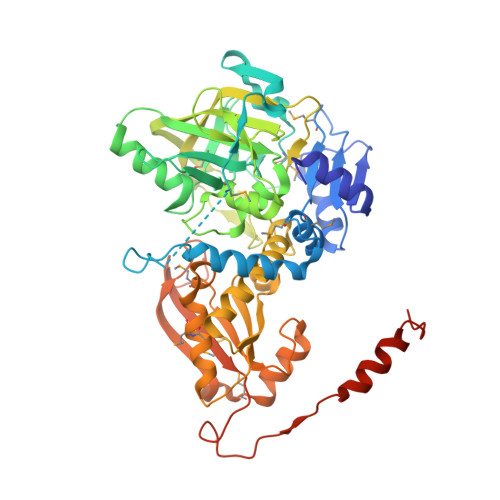
Crystal Structure of 3-octaprenyl-4-hydroxybenzoate decarboxylase (UbiD) from Escherichia coli, Northeast Structural Genomics Target ER459.
Zhou, W., Forouhar, F., Seetharaman, J., Fang, Y., Xiao, R., Cunningham, K., Ma, L.-C., Chen, C.X., Acton, T.B., Montelione, G.T., Hunt, J.F., Tong, L., Northeast Structural Genomics Consortium (NESG)To be published.