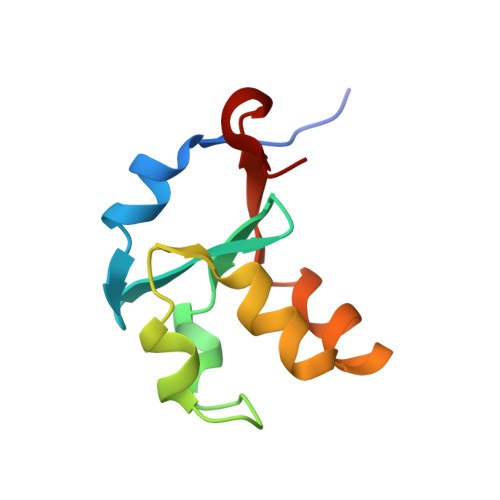
Comparison of cytochromes b(5) from insects and vertebrates.
Wang, L., Cowley, A.B., Terzyan, S., Zhang, X., Benson, D.R.(2007) Proteins 67: 293-304
- PubMed: 17299762 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21250
- Primary Citation Related Structures:
2IBJ - PubMed Abstract:
We report a 1.55 A X-ray crystal structure of the heme-binding domain of cytochrome b(5) from Musca domestica (house fly; HF b(5)), and compare it with previously published structures of the heme-binding domains of bovine microsomal cytochrome b(5) (bMc b(5)) and rat outer mitochondrial membrane cytochrome b(5) (rOM b(5)). The structural comparison was done in the context of amino acid sequences of all known homologues of the proteins under study. We show that insect b(5)s contain an extended hydrophobic patch at the base of the heme binding pocket, similar to the one previously shown to stabilize mammalian OM b(5)s relative to their Mc counterparts. The hydrophobic patch in insects includes a residue with a bulky hydrophobic side chain at position 71 (Met). Replacing Met71 in HF b(5) with Ser, the corresponding residue in all known mammalian Mc b(5)s, is found to substantially destabilize the holoprotein. The destabilization is a consequence of two related factors: (1) a large decrease in apoprotein stability and (2) extension of conformational disruption in the apoprotein beyond the empty heme binding pocket (core 1) and into the heme-independent folding core (core 2). Analogous changes have previously been shown to accompany replacement of Leu71 in rOM b(5) with Ser. That the stabilizing role of Met71 in HF b(5) is manifested primarily in the apo state is highlighted by the fact that its crystallographic Calpha B factor is modestly larger than that of Ser71 in bMc b(5), indicating that it slightly destabilizes local polypeptide conformation when heme is in its binding pocket. Finally, we show that the final unit of secondary structure in the cytochrome b(5) heme-binding domain, a 3(10) helix known as alpha6, differs substantially in length and packing interactions not only for different protein isoforms but also for given isoforms from different species.
- Department of Chemistry, University of Kansas, Lawrence, Kansas 66045, USA.
Organizational Affiliation: