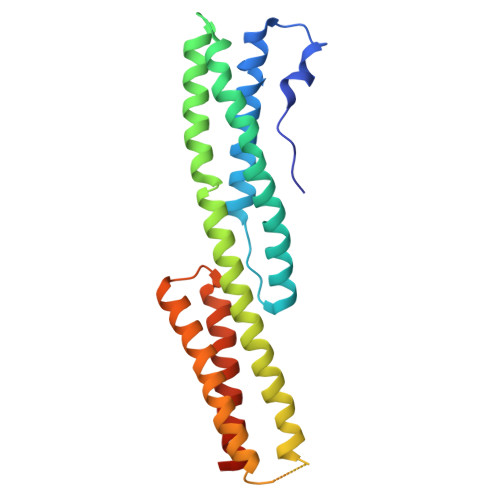
Structural analysis of the plakin domain of bullous pemphigoid antigen1 (BPAG1) suggests that plakins are members of the spectrin superfamily.
Jefferson, J.J., Ciatto, C., Shapiro, L., Liem, R.K.(2007) J Mol Biology 366: 244-257
- PubMed: 17161423 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2006.11.036
- Primary Citation Related Structures:
2IAK - PubMed Abstract:
Bullous pemphigoid antigen 1 (BPAG1) is a member of the plakin family of proteins. The plakins are multi-domain proteins that have been shown to interact with microtubules, actin filaments and intermediate filaments, as well as proteins found in cellular junctions. These interactions are mediated through different domains on the plakins. The interactions between plakins and components of specialized cell junctions such as desmosomes and hemidesmosomes are mediated through the so-called plakin domain, which is a common feature of the plakins. We report the crystal structure of a stable fragment from BPAG1, residues 226-448, defined by limited proteolysis of the whole plakin domain. The structure, determined by single-wavelength anomalous diffraction phasing from a selenomethionine-substituted crystal at 3.0 A resolution, reveals a tandem pair of triple helical bundles closely related to spectrin repeats. Based on this structure and analysis of sequence conservation, we propose that the architecture of plakin domains is defined by two pairs of spectrin repeats interrupted by a putative Src-Homology 3 (SH3) domain.
- Department of Pathology and Cell Biology, Columbia University College of Physicians and Surgeons, New York, NY 10032, USA.
Organizational Affiliation: