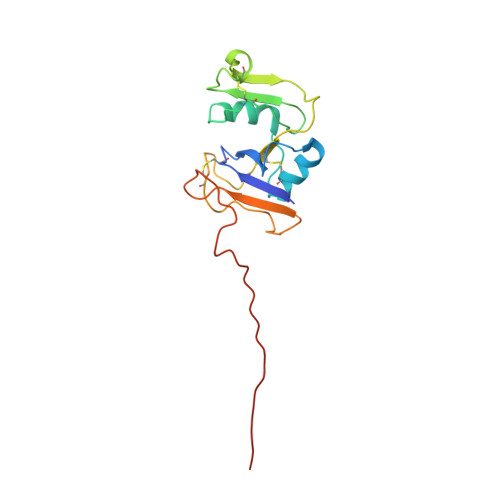
Ligand-induced Structural Changes of the CD44 Hyaluronan-binding Domain Revealed by NMR
Takeda, M., Ogino, S., Umemoto, R., Sakakura, M., Kajiwara, M., Sugahara, K.N., Hayasaka, H., Miyasaka, M., Terasawa, H., Shimada, I.(2006) J Biological Chem 281: 40089-40095
- PubMed: 17085435 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M608425200
- Primary Citation Related Structures:
2I83 - PubMed Abstract:
CD44, a major cell surface receptor for hyaluronan (HA), contains a functional domain responsible for HA binding at its N terminus (residues 21-178). Accumulating evidence indicates that proteolytic cleavage of CD44 in its extracellular region (residues 21-268) leads to enhanced tumor cell migration and invasion. Hence, understanding the mechanisms underlying the CD44 proteolytic cleavage is important for understanding the mechanism of CD44-mediated tumor progression. Here we present the NMR structure of the HA-binding domain of CD44 in its HA-bound state. The structure is composed of the Link module (residues 32-124) and an extended lobe (residues 21-31 and 125-152). Interestingly, a comparison of its unbound and HA-bound structures revealed that rearrangement of the beta-strands in the extended lobe (residues 143-148) and disorder of the structure in the following C-terminal region (residues 153-169) occurred upon HA binding, which is consistent with the results of trypsin proteolysis studies of the CD44 HA-binding domain. The order-to-disorder transition of the C-terminal region by HA binding may be involved in the CD44-mediated cell migration.
- Graduate School of Pharmaceutical Sciences, University of Tokyo, Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: