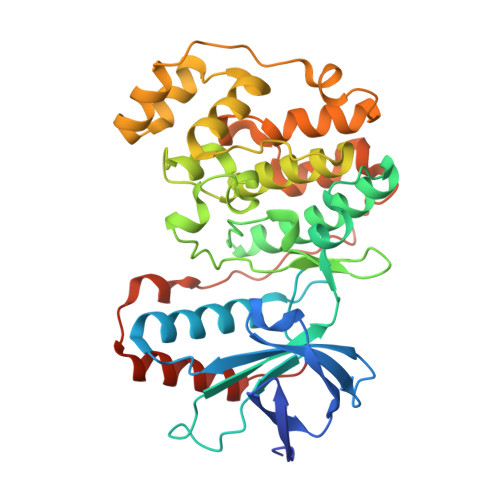
p38 MAP Kinase Inhibitors Part 6: 2-Arylpyridazin-3-ones as templates for inhibitor design.
Natarajan, S.R., Heller, S.T., Nam, K., Singh, S.B., Scapin, G., Patel, S., Fitzgerald, C.E., Thompson, J.E., O'Keefe, S.J.(2006) Bioorg Med Chem Lett 16: 5809-5813
- PubMed: 16945533 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.08.074
- Primary Citation Related Structures:
2I0H - PubMed Abstract:
p38 inhibitors based on 3,4-dihydropyrido[4,3-d]pyrimidazin-2-one template were synthesized and their SAR explored. Benchmark compounds 30, 35, and 36 were found to be potent against the enzyme. Crystal structure of p38 in complex with 30 indicated a key pi-stacking interaction with the pendant tyrosine residue-35 in the glycine-rich loop.
- Department of Medicinal Chemistry, Merck Research Laboratories, PO Box 2000, Rahway, NJ 07065, USA. ravi_natarajan@merck.com
Organizational Affiliation: