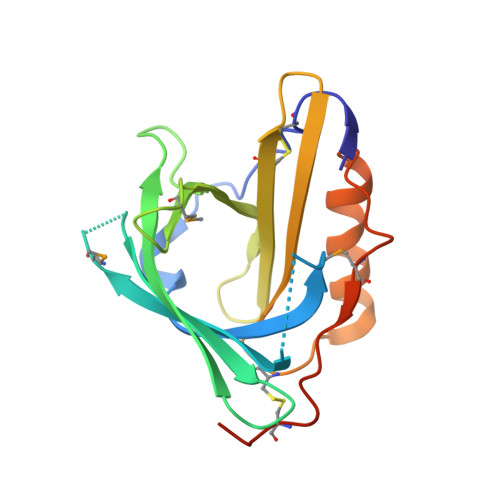
Structural insight into the dual ligand specificity and mode of high density lipoprotein association of apolipoprotein d.
Eichinger, A., Nasreen, A., Kim, H.J., Skerra, A.(2007) J Biological Chem 282: 31068-31075
- PubMed: 17699160 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M703552200
- Primary Citation Related Structures:
2HZQ, 2HZR - PubMed Abstract:
Human apolipoprotein D (ApoD) occurs in plasma associated with high density lipoprotein. Apart from the involvement in lipid metabolism, its binding activity for progesterone and arachidonic acid plays a role in cancer development and neurological diseases. The crystal structures of free ApoD and its complex with progesterone were determined at 1.8A resolution and reveal a lipocalin fold. The narrow, mainly uncharged pocket within the typical beta-barrel accommodates progesterone with its acetyl side chain oriented toward the bottom. The cavity adopts essentially the same shape in the absence of progesterone and allows complexation of arachidonic acid as another cognate ligand. Three of the four extended loops at the open end of the beta-barrel expose hydrophobic side chains, which is an unusual feature for lipocalins and probably effects association with the high density lipoprotein particle by mediating insertion into the lipid phase. This mechanism is in line with an unpaired Cys residue in the same surface region that can form a disulfide cross-link with apolipoprotein A-II.
- Lehrstuhl für Biologische Chemie, Technische Universität München, 85350 Freising-Weihenstephan, Germany.
Organizational Affiliation: