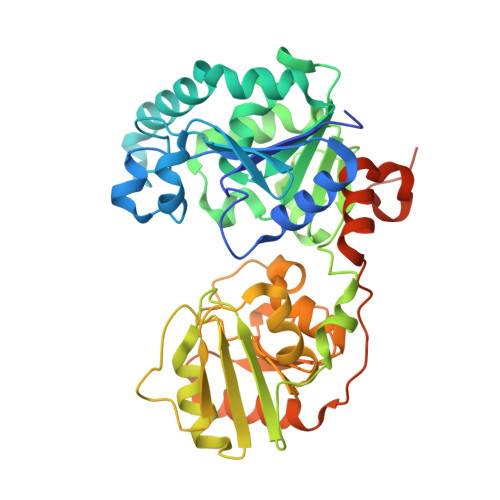
Structure and mechanism of GumK, a membrane-associated glucuronosyltransferase.
Barreras, M., Salinas, S.R., Abdian, P.L., Kampel, M.A., Ielpi, L.(2008) J Biological Chem 283: 25027-25035
- PubMed: 18596046
- DOI: https://doi.org/10.1074/jbc.M801227200
- Primary Citation Related Structures:
2HY7, 2Q6V, 3CUY, 3CV3 - PubMed Abstract:
Xanthomonas campestris GumK (beta-1,2-glucuronosyltransferase) is a 44-kDa membrane-associated protein that is involved in the biosynthesis of xanthan, an exopolysaccharide crucial for this bacterium's phytopathogenicity. Xanthan also has many important industrial applications. The GumK enzyme is the founding member of the glycosyltransferase family 70 of carbohydrate-active enzymes, which is composed of bacterial glycosyltransferases involved in exopolysaccharide synthesis. No x-ray structures have been reported for this family. To better understand the mechanism of action of the bacterial glycosyltransferases in this family, the x-ray crystal structure of apo-GumK was solved at 1.9 angstroms resolution. The enzyme has two well defined Rossmann domains with a catalytic cleft between them, which is a typical feature of the glycosyltransferase B superfamily. Additionally, the crystal structure of GumK complexed with UDP was solved at 2.28 angstroms resolution. We identified a number of catalytically important residues, including Asp157, which serves as the general base in the transfer reaction. Residues Met231, Met273, Glu272, Tyr292, Met306, Lys307, and Gln310 interact with UDP, and mutation of these residues affected protein activity both in vitro and in vivo. The biological and structural data reported here shed light on the molecular basis for donor and acceptor selectivity in this glycosyltransferase family. These results also provide a rationale to obtain new polysaccharides by varying residues in the conserved alpha/beta/alpha structural motif of GumK.
- Laboratory of Bacterial Genetics, Fundación Instituto Leloir, IIBBA-Consejo Nacional de Investigaciones Científicas y Técnicas, Buenos Aires C1405BWE, Argentina.
Organizational Affiliation: