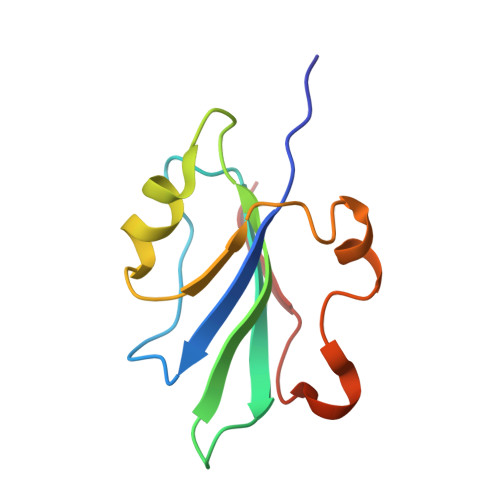
Solution structure of the endonuclease domain from the master replication initiator protein of the nanovirus faba bean necrotic yellows virus and comparison with the corresponding geminivirus and circovirus structures
Vega-Rocha, S., Gronenborn, B., Gronenborn, A.M., Campos-Olivas, R.(2007) Biochemistry 46: 6201-6212
- PubMed: 17472345
- DOI: https://doi.org/10.1021/bi700159q
- Primary Citation Related Structures:
2HWT - PubMed Abstract:
Nanoviruses are a family of plant viruses that possess a genome of multiple circular single-stranded DNA (ssDNA) components and are strikingly similar in their replication mode to the plant geminiviruses and to the circoviruses that infect birds or mammals. These viruses multiply by rolling circle replication using virus-encoded multifunctional replication initiator proteins (Rep proteins) that catalyze the initiation of replication on a double-stranded DNA (dsDNA) intermediate and the resolution of the ssDNA into circles. Here we report the solution NMR three-dimensional structure of the endonuclease domain from the master Rep (M-Rep) protein of faba bean necrotic yellows virus (FBNYV), a representative of the nanoviruses. The domain comprises amino acids 2-95 (M-Rep2-95), and its global fold is similar to those previously described for the gemini- and circovirus Rep endonuclease domains, consisting of a central 5-stranded antiparallel beta-sheet covered on one side by an alpha-helix and irregular loops and on the other, more open side of the domain, by an alpha-helix containing the catalytic tyrosine residue (the catalytic helix). Longer domain constructs extending to amino acids 117 and 124 were also characterized. They contain an additional alpha-helix, are monomeric, and exhibit catalytic activity indistinguishable from that of M-Rep2-95. The binding site for the catalytic metal was identified by paramagnetic broadening and maps to residues on the exposed face of the central beta-sheet. A comparison with the previously determined Rep endonuclease domain structures of tomato yellow leaf curl Sardinia virus (TYLCSV), a geminivirus, and that of porcine circovirus type 2 (PCV2) Rep allows the identification of a positively charged surface that is most likely involved in dsDNA binding, and reveals common features shared by all endonuclease domains of nanovirus, geminivirus, and circovirus Rep proteins.
- Structural and Computational Biology Program, Spanish National Cancer Center (CNIO), Madrid 28029, Spain.
Organizational Affiliation: