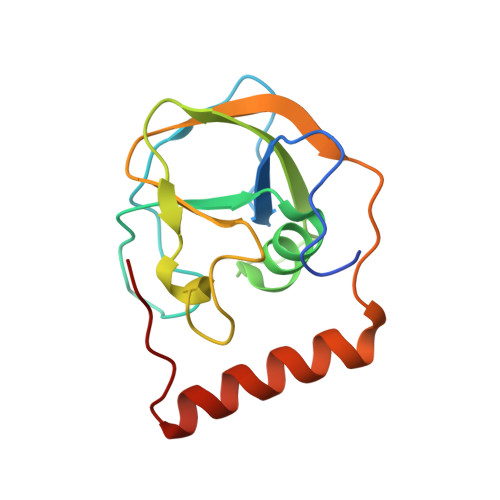
Rifamycin antibiotic resistance by ADP-ribosylation: Structure and diversity of Arr.
Baysarowich, J., Koteva, K., Hughes, D.W., Ejim, L., Griffiths, E., Zhang, K., Junop, M., Wright, G.D.(2008) Proc Natl Acad Sci U S A 105: 4886-4891
- PubMed: 18349144
- DOI: https://doi.org/10.1073/pnas.0711939105
- Primary Citation Related Structures:
2HW2 - PubMed Abstract:
The rifamycin antibiotic rifampin is important for the treatment of tuberculosis and infections caused by multidrug-resistant Staphylococcus aureus. Recent iterations of the rifampin core structure have resulted in new drugs and drug candidates for the treatment of a much broader range of infectious diseases. This expanded use of rifamycin antibiotics has the potential to select for increased resistance. One poorly characterized mechanism of resistance is through Arr enzymes that catalyze ADP-ribosylation of rifamycins. We find that genes encoding predicted Arr enzymes are widely distributed in the genomes of pathogenic and nonpathogenic bacteria. Biochemical analysis of three representative Arr enzymes from environmental and pathogenic bacterial sources shows that these have equally efficient drug resistance capacity in vitro and in vivo. The 3D structure of one of these orthologues from Mycobacterium smegmatis was determined and reveals structural homology with ADP-ribosyltransferases important in eukaryotic biology, including poly(ADP-ribose) polymerases (PARPs) and bacterial toxins, despite no significant amino acid sequence homology with these proteins. This work highlights the extent of the rifamycin resistome in microbial genera with the potential to negatively impact the expanded use of this class of antibiotic.
- M.G. DeGroote Institute for Infectious Disease Research, Department of Biochemistry and Biomedical Sciences, McMaster University, Hamilton, ON, Canada L8N 3Z5.
Organizational Affiliation: