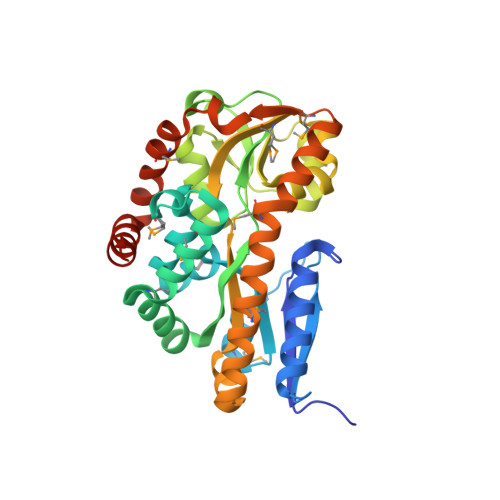
Structural analysis of a periplasmic binding protein in the tripartite ATP-independent transporter family reveals a tetrameric assembly that may have a role in ligand transport.
Cuneo, M.J., Changela, A., Miklos, A.E., Beese, L.S., Krueger, J.K., Hellinga, H.W.(2008) J Biological Chem 283: 32812-32820
- PubMed: 18723845 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M803595200
- Primary Citation Related Structures:
2HPG - PubMed Abstract:
Several bacterial solute transport mechanisms involve members of the periplasmic binding protein (PBP) superfamily that bind and deliver ligand to integral membrane transport proteins in the ATP-binding cassette, tripartite tricarboxylate transporter, or tripartite ATP-independent (TRAP) families. PBPs involved in ATP-binding cassette transport systems have been well characterized, but only a few PBPs involved in TRAP transport have been studied. We have measured the thermal stability, determined the oligomerization state by small angle x-ray scattering, and solved the x-ray crystal structure to 1.9 A resolution of a TRAP-PBP (open reading frame tm0322) from the hyperthermophilic bacterium Thermotoga maritima (TM0322). The overall fold of TM0322 is similar to other TRAP transport related PBPs, although the structural similarity of backbone atoms (2.5-3.1 A root mean square deviation) is unusually low for PBPs within the same group. Individual monomers within the tetrameric asymmetric unit of TM0322 exhibit high root mean square deviation (0.9 A) to each other as a consequence of conformational heterogeneity in their binding pockets. The gel filtration elution profile and the small angle x-ray scattering analysis indicate that TM0322 assembles as dimers in solution that in turn assemble into a dimer of dimers in the crystallographic asymmetric unit. Tetramerization has been previously observed in another TRAP-PBP (the Rhodobacter sphaeroides alpha-keto acid-binding protein) where quaternary structure formation is postulated to be an important requisite for the transmembrane transport process.
- Department of Biochemistry, Duke University, Medical Center, Durham, North Carolina 27710, USA.
Organizational Affiliation: