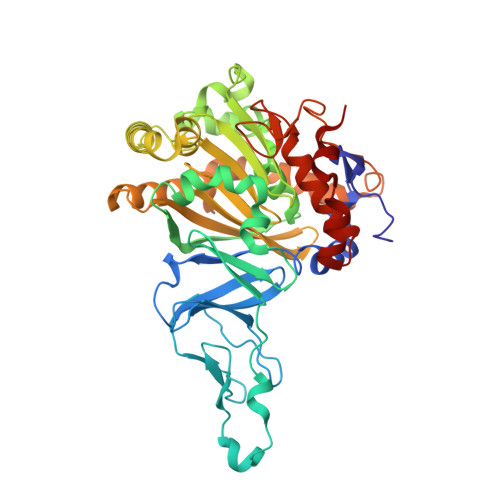
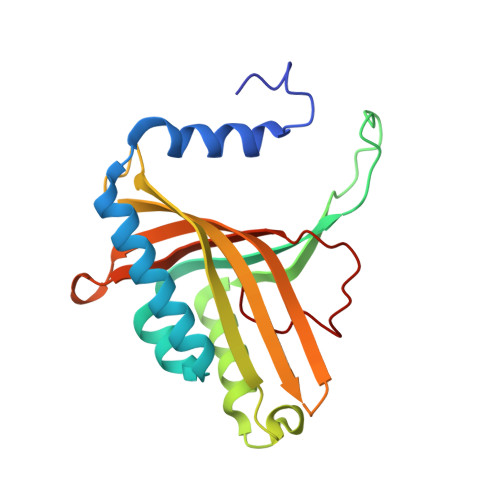
Structural basis for regioselectivity and stereoselectivity of product formation by naphthalene 1,2-dioxygenase.
Ferraro, D.J., Okerlund, A.L., Mowers, J.C., Ramaswamy, S.(2006) J Bacteriol 188: 6986-6994
- PubMed: 16980501 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00707-06
- Primary Citation Related Structures:
2HMJ, 2HMK, 2HML, 2HMM, 2HMN, 2HMO - PubMed Abstract:
Rieske oxygenase (RO) systems are two- and three-component enzyme systems that catalyze the formation of cis-dihydrodiols from aromatic substrates. Degradation of pollutants in contaminated soil and generation of chiral synthons have been the major foci of RO research. Substrate specificity and product regio- and stereoselectivity have been shown to vary between individual ROs. While directed evolution methods for altering RO function have been successful in the past, rational engineering of these enzymes still poses a challenge due to the lack of structural understanding. Here we examine the structural changes induced by mutation of Phe-352 in naphthalene 1,2-dioxygenase from Pseudomonas sp. strain NCIB 9816-4 (NDO-O(9816-4)). Structures of the Phe-352-Val mutant in native form and in complex with phenanthrene and anthracene, along with those of wild-type NDO-O(9816-4) in complex with phenanthrene, anthracene, and 3-nitrotoluene, are presented. Phenanthrene was shown to bind in a different orientation in the Phe-352-Val mutant active site from that in the wild type, while anthracene was found to bind in similar positions in both enzymes. Two orientations of 3-nitrotoluene were observed, i.e., a productive and a nonproductive orientation. These orientations help explain why NDO-O(9816-4) forms different products from 3-nitrotoluene than those made from nitrobenzene dioxygenase. Comparison of these structures among themselves and with other known ROs bound to substrates reveals that the orientation of substrate binding at the active site is the primary determinant of product regio- and stereoselectivity.
- Department of Biochemistry, Roy J. and Lucille A. Carver College of Medicine, University of Iowa, 51 Newton Road, 4-403 BSB, Iowa City, IA 52242, USA.
Organizational Affiliation: