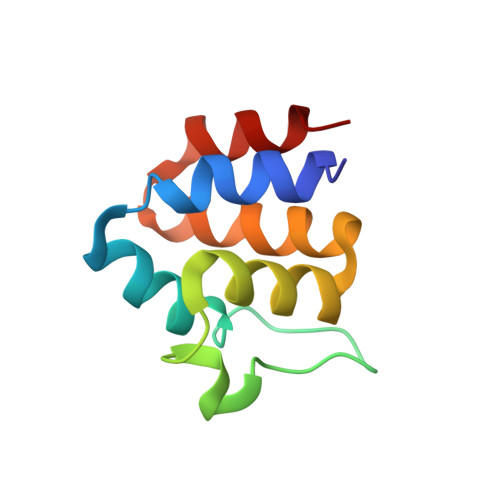
Structure and dynamics of ASC2, a pyrin domain-only protein that regulates inflammatory signaling
Natarajan, A., Ghose, R., Hill, J.M.(2006) J Biological Chem 281: 31863-31875
- PubMed: 16905547
- DOI: https://doi.org/10.1074/jbc.M605458200
- Primary Citation of Related Structures:
2HM2 - PubMed Abstract:
Pyrin domain (PYD)-containing proteins are key components of pathways that regulate inflammation, apoptosis, and cytokine processing. Their importance is further evidenced by the consequences of mutations in these proteins that give rise to autoimmune and hyperinflammatory syndromes. PYDs, like other members of the death domain (DD) superfamily, are postulated to mediate homotypic interactions that assemble and regulate the activity of signaling complexes. However, PYDs are presently the least well characterized of all four DD subfamilies. Here we report the three-dimensional structure and dynamic properties of ASC2, a PYD-only protein that functions as a modulator of multidomain PYD-containing proteins involved in NF-kappaB and caspase-1 activation. ASC2 adopts a six-helix bundle structure with a prominent loop, comprising 13 amino acid residues, between helices two and three. This loop represents a divergent feature of PYDs from other domains with the DD fold. Detailed analysis of backbone 15N NMR relaxation data using both the Lipari-Szabo model-free and reduced spectral density function formalisms revealed no evidence of contiguous stretches of polypeptide chain with dramatically increased internal motion, except at the extreme N and C termini. Some mobility in the fast, picosecond to nanosecond timescale, was seen in helix 3 and the preceding alpha2-alpha3 loop, in stark contrast to the complete disorder seen in the corresponding region of the NALP1 PYD. Our results suggest that extensive conformational flexibility in helix 3 and the alpha2-alpha3 loop is not a general feature of pyrin domains. Further, a transition from complete disorder to order of the alpha2-alpha3 loop upon binding, as suggested for NALP1, is unlikely to be a common attribute of pyrin domain interactions.
- Department of Chemistry, City College of the City University of New York, New York, New York 10031, USA.
Organizational Affiliation: