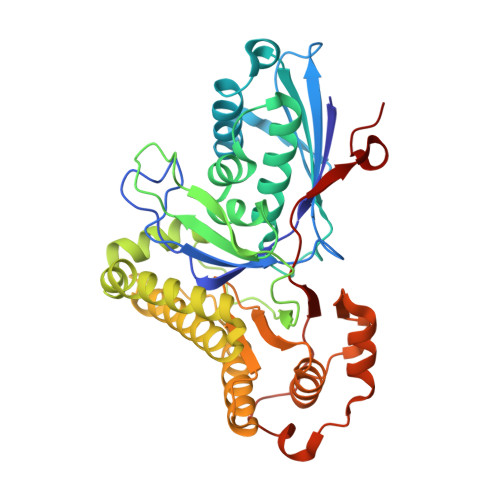
Crystal Structures of Trypanosoma brucei and Staphylococcus aureus Mevalonate Diphosphate Decarboxylase Inform on the Determinants of Specificity and Reactivity
Byres, E., Alphey, M.S., Smith, T.K., Hunter, W.N.(2007) J Mol Biology 371: 540-553
- PubMed: 17583736 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2007.05.094
- Primary Citation Related Structures:
2HK2, 2HK3, 2HKE - PubMed Abstract:
Mevalonate diphosphate decarboxylase (MDD) catalyzes the ATP-dependent decarboxylation of mevalonate 5-diphosphate (MDP) to form isopentenyl pyrophosphate, a ubiquitous precursor for isoprenoid biosynthesis. MDD is a poorly understood component of this important metabolic pathway. Complementation of a temperature-sensitive yeast mutant by the putative mdd genes of Trypanosoma brucei and Staphylococcus aureus provides proof-of-function. Crystal structures of MDD from T. brucei (TbMDD, at 1.8 A resolution) and S. aureus (SaMDD, in two distinct crystal forms, each diffracting to 2.3 A resolution) have been determined. Gel-filtration chromatography and analytical ultracentrifugation experiments indicate that TbMDD is predominantly monomeric in solution while SaMDD is dimeric. The new crystal structures and comparison with that of the yeast Saccharomyces cerevisiae enzyme (ScMDD) reveal the structural basis for this variance in quaternary structure. The presence of an ordered sulfate in the structure of TbMDD reveals for the first time details of a ligand binding in the MDD active site and, in conjunction with well-ordered water molecules, comparisons with the related enzyme mevalonate kinase, structural and biochemical data derived on ScMDD and SaMDD, allows us to model a ternary complex with MDP and ATP. This model facilitates discussion of the molecular determinants of substrate recognition and contributions made by specific residues to the enzyme mechanism.
- Division of Biological Chemistry and Molecular Microbiology, College of Life Sciences, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: