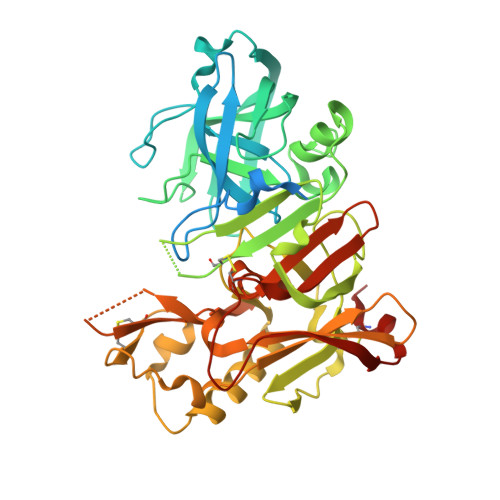
Design of potent inhibitors of human beta-secretase. Part 1.
Freskos, J.N., Fobian, Y.M., Benson, T.E., Bienkowski, M.J., Brown, D.L., Emmons, T.L., Heintz, R., Laborde, A., McDonald, J.J., Mischke, B.V., Molyneaux, J.M., Moon, J.B., Mullins, P.B., Bryan Prince, D., Paddock, D.J., Tomasselli, A.G., Winterrowd, G.(2007) Bioorg Med Chem Lett 17: 73-77
- PubMed: 17046251 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2006.09.092
- Primary Citation Related Structures:
2HIZ - PubMed Abstract:
We describe a novel series of potent inhibitors of human beta-secretase. These compounds possess the hydroxyethyl amine transition state isostere. A 2.5A crystal structure of inhibitor 32 bound to BACE is provided.
- Pfizer Inc., 700N. Chesterfield Pkwy., St. Louis, MO 63198, USA. john.n.freskos@pfizer.com
Organizational Affiliation: