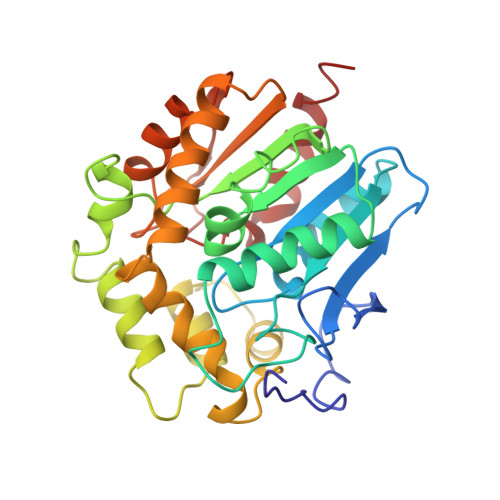
Crystal structure of haloalkane dehalogenase: an enzyme to detoxify halogenated alkanes.
Franken, S.M., Rozeboom, H.J., Kalk, K.H., Dijkstra, B.W.(1991) EMBO J 10: 1297-1302
- PubMed: 2026135 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1991.tb07647.x
- Primary Citation Related Structures:
2HAD - PubMed Abstract:
Haloalkane dehalogenase from Xanthobacter autotrophicus GJ10 converts 1-haloalkanes to the corresponding alcohols and halide ions with water as the sole cosubstrate and without any need for oxygen or cofactors. The three-dimensional structure has been determined by multiple isomorphous replacement techniques using three heavy atom derivatives. The structure has been refined at 2.4 A resolution to an R-factor of 17.9%. The monomeric enzyme is a spherical molecule and is composed to two domains: domain I has an alpha/beta type structure with a central eight-stranded mainly parallel beta-sheet. Domain II lies like a cap on top of domain I and consists of alpha-helices connected by loops. Except for the cap domain the structure resembles that of the dienelactone hydrolase in spite of any significant sequence homology. The putative active site is completely buried in an internal hydrophobic cavity which is located between the two domains. From the analysis of the structure it is suggested that Asp124 is the nucleophilic residue essential for the catalysis. It interacts with His289 which is hydrogen-bonded to Asp260.
- Laboratory of Chemical Physics, Department of Chemistry, Groningen, The Netherlands.
Organizational Affiliation: