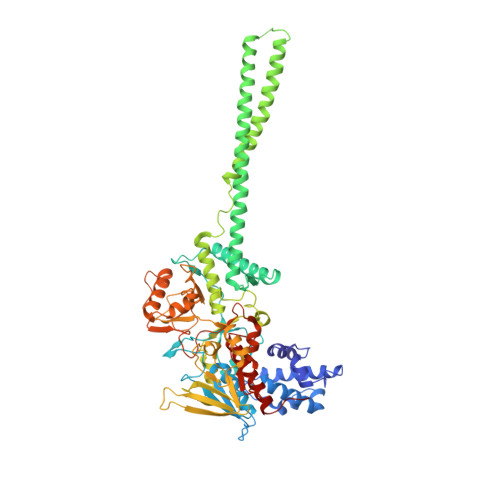
Crystal structure and mechanism of human lysine-specific demethylase-1.
Stavropoulos, P., Blobel, G., Hoelz, A.(2006) Nat Struct Mol Biol 13: 626-632
- PubMed: 16799558 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb1113
- Primary Citation Related Structures:
2H94 - PubMed Abstract:
The reversible methylation of specific lysine residues in histone tails is crucial in epigenetic gene regulation. LSD1, the first known lysine-specific demethylase, selectively removes monomethyl and dimethyl, but not trimethyl modifications of Lys4 or Lys9 of histone-3. Here, we present the crystal structure of LSD1 at 2.9-A resolution. LSD1 forms a highly asymmetric, closely packed domain structure from which a long helical 'tower' domain protrudes. The active site cavity is spacious enough to accommodate several residues of the histone tail substrate, but does not appear capable of recognizing the different methylation states of the substrate lysine. This supports the hypothesis that trimethylated lysine is chemically rather than sterically discriminated. We present a biochemical analysis of LSD1 mutants that identifies crucial residues in the active site cavity and shows the importance of the SWIRM and tower domains for catalysis.
- Laboratory of Cell Biology, Howard Hughes Medical Institute, The Rockefeller University, 1230 York Avenue, New York, New York 10021, USA.
Organizational Affiliation: