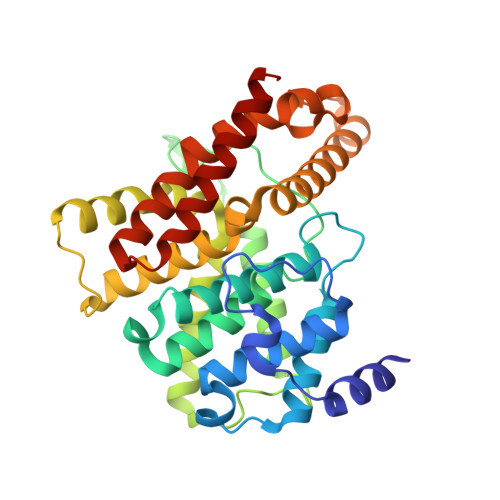
Multiple Conformations of Phosphodiesterase-5: Implications for enzyme function and drug development
Wang, H., Liu, Y., Huai, Q., Cai, J., Zoraghi, R., Francis, S.H., Corbin, J.D., Robinson, H., Xin, Z., Lin, G., Ke, H.(2006) J Biological Chem 281: 21469-21479
- PubMed: 16735511 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M512527200
- Primary Citation Related Structures:
2H40, 2H42, 2H44 - PubMed Abstract:
Phosphodiesterase-5 (PDE5) is the target for sildenafil, vardenafil, and tadalafil, which are drugs for treatment of erectile dysfunction and pulmonary hypertension. We report here the crystal structures of a fully active catalytic domain of unliganded PDE5A1 and its complexes with sildenafil or icarisid II. These structures together with the PDE5A1-isobutyl-1-methylxanthine complex show that the H-loop (residues 660-683) at the active site of PDE5A1 has four different conformations and migrates 7-35A upon inhibitor binding. In addition, the conformation of sildenafil reported herein differs significantly from those in the previous structures of chimerically hybridized or almost inactive PDE5. Mutagenesis and kinetic analyses confirm that the H-loop is particularly important for substrate recognition and that invariant Gly(659), which immediately precedes the H-loop, is critical for optimal substrate affinity and catalytic activity.
- Department of Biochemistry and Biophysics and Lineberger Comprehensive Cancer Center, University of North Carolina, Chapel Hill, North Carolina 27599-7260.
Organizational Affiliation: