Structural Basis for Nucleic Acid and Toxin Recognition of the Bacterial Antitoxin CcdA
Madl, T., Van Melderen, L., Mine, N., Respondek, M., Oberer, M., Keller, W., Khatai, L., Zangger, K.(2006) J Mol Biology 364: 170-185
- PubMed: 17007877 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.08.082
- Primary Citation Related Structures:
2ADL, 2ADN, 2H3A, 2H3C - PubMed Abstract:
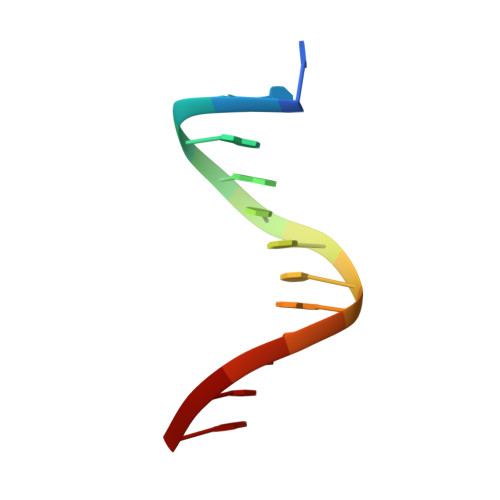
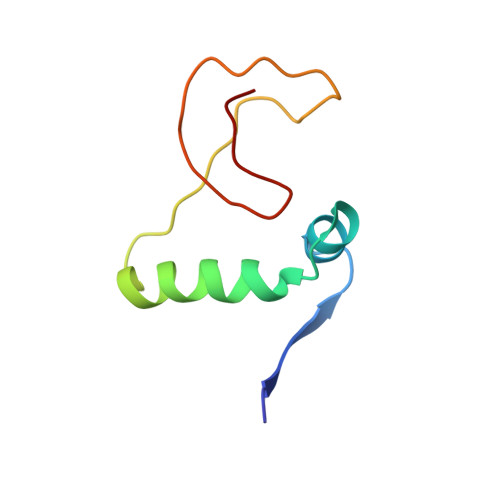
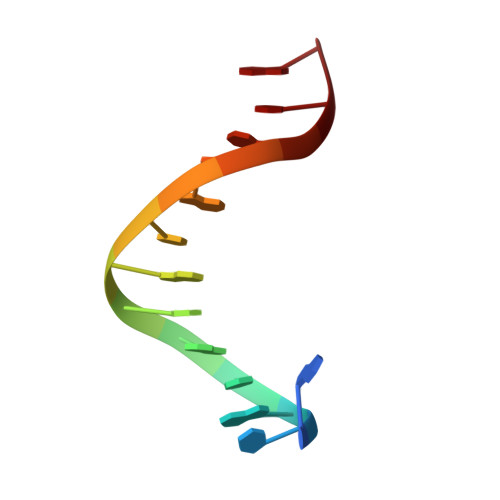
Toxin-antitoxin systems are highly abundant in plasmids and bacterial chromosomes. They ensure plasmid maintenance by killing bacteria that have lost the plasmid. Their expression is autoregulated at the level of transcription. Here, we present the solution structure of CcdA, the antitoxin of the ccd system, as a free protein (16.7 kDa) and in complex with its cognate DNA (25.3 kDa). CcdA is composed of two distinct and independent domains: the N-terminal domain, responsible for DNA binding, which establishes a new family of the ribbon-helix-helix fold and the C-terminal region, which is responsible for the interaction with the toxin CcdB. The C-terminal domain is intrinsically unstructured and forms a tight complex with the toxin. We show that CcdA specifically recognizes a 6 bp palindromic DNA sequence within the operator-promoter (OP) region of the ccd operon and binds to DNA by insertion of the positively charged N-terminal beta-sheet into the major groove. The binding of up to three CcdA dimers to a 33mer DNA of its operator-promoter region was studied by NMR spectroscopy, isothermal titration calorimetry and single point mutation. The highly flexible C-terminal region of free CcdA explains its susceptibility to proteolysis by the Lon ATP-dependent protease.
- Institute of Chemistry, University of Graz, Graz 8010, Austria.
Organizational Affiliation: