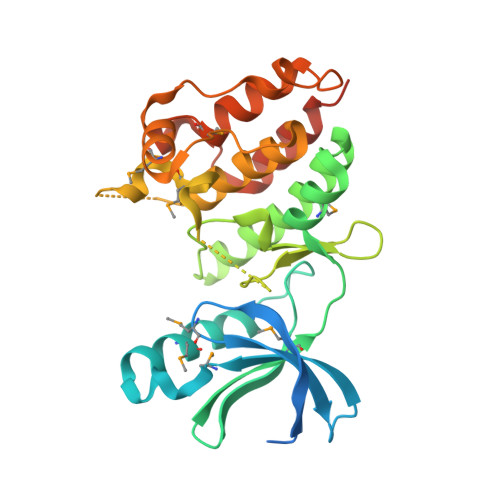
A Conserved Dimer and Global Conformational Changes in the Structure of apo-PknE Ser/Thr Protein Kinase from Mycobacterium tuberculosis.
Gay, L.M., Ng, H.L., Alber, T.(2006) J Mol Biology 360: 409-420
- PubMed: 16762364 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.05.015
- Primary Citation Related Structures:
2H34 - PubMed Abstract:
The "eukaryotic-like" receptor Ser/Thr protein kinases (STPKs) are candidates for the sensors that mediate environmental adaptations of Mycobacterium tuberculosis (Mtb). To define the mechanisms of regulation and substrate recognition, we determined the crystal structure of the ligand-free, activated kinase domain (KD) of the Mtb STPK, PknE. Remarkably, the PknE KD formed a dimer similar to that first observed in the structure of the ATPgammaS complex of the Mtb paralog, PknB. This structural similarity, which occurs despite little sequence conservation between the PknB and PknE dimer interfaces, supports the idea that dimerization regulates the Mtb receptor STPKs. Insertion of the DFG motif into the ATP-binding site and other conformational differences compared the ATPgammaS:PknB complex suggest that apo-PknE is not pre-organized to bind nucleotides. This structure may represent an inactive conformation stabilized by dimerization or, alternatively, an active conformation that reveals shifts that mediate nucleotide exchange and order substrate binding.
- Department of Molecular and Cell Biology, University of California-Berkeley, Berkeley, CA 94720-3206, USA.
Organizational Affiliation: