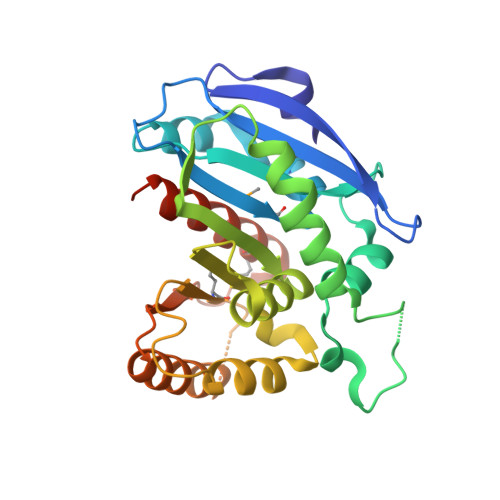
Structural Characterization of Enterobactin Hydrolase IroE.
Larsen, N.A., Lin, H., Wei, R., Fischbach, M.A., Walsh, C.T.(2006) Biochemistry 45: 10184-10190
- PubMed: 16922493
- DOI: https://doi.org/10.1021/bi060950i
- Primary Citation of Related Structures:
2GZR, 2GZS - PubMed Abstract:
The proliferation of many pathogenic bacteria is limited by the scarcity of soluble iron in their environment. Many of these bacteria scavenge iron by synthesizing and exporting small molecule siderophores that chelate iron. Iron-bound siderophores are subsequently imported for metabolic processing. Three related serine hydrolases have been characterized biochemically in this pathway: Fes, IroD, and IroE. Here, we report the crystal structure of IroE from uropathogenic Escherichia coli CFT073. The native structure and a complex with diisopropyl fluorophosphonate (DFP, a potent serine hydrolase inhibitor) were determined at 2.3 and 1.4 A resolution, respectively. IroE has the typical alpha/beta-hydrolase fold with an atypical catalytic dyad composed of Ser 189 and His 287. Mutation of either residue was detrimental to catalysis. In addition, rather than the typical oxyanion hole composed of backbone amides, IroE employs the atypical guanidinium moiety of Arg 130. Asp 90 anchors Arg 130 in the active site, and mutation of either residue was likewise detrimental to catalysis. We also compare the structure of IroE to the structure of Fes from Shigella flexneri (PDB entry 2B20). Both enzymes have similar active sites, but Fes has an additional amino-terminal lid domain. These lid domains are proposed to confer specificity to these related hydrolases.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, 240 Longwood Avenue, Boston, Massachusetts 02115-5702, USA.
Organizational Affiliation: