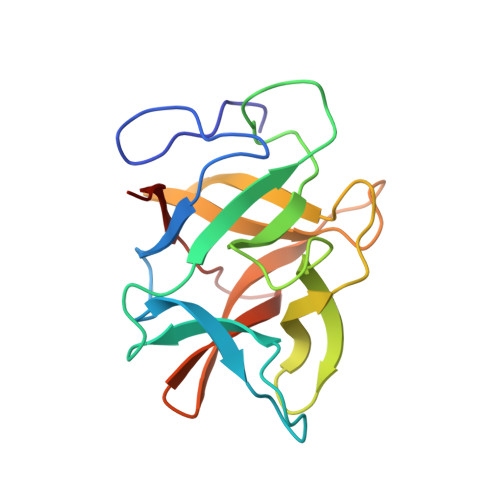
Crystal structure of a novel cysteinless plant Kunitz-type protease inhibitor.
Hansen, D., Macedo-Ribeiro, S., Verissimo, P., Yoo Im, S., Sampaio, M.U., Oliva, M.L.(2007) Biochem Biophys Res Commun 360: 735-740
- PubMed: 17631863 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2007.06.144
- Primary Citation Related Structures:
2GZB - PubMed Abstract:
Bauhinia bauhinioides Cruzipain Inhibitor (BbCI) is a cysteine protease inhibitor highly homologous to plant Kunitz-type inhibitors. However, in contrast to classical Kunitz family inhibitors it lacks cysteine residues and therefore disulfide bridges. BbCI is also distinct in the ability to inactivate enzymes belonging to two different classes, cysteine and serine proteases. Besides inhibiting the cysteine protease cruzipain, BbCI also inhibits cathepsin L and the serine proteases HNE (human neutrophil elastase) and PPE (porcine pancreatic elastase). Monoclinic crystals of the recombinant inhibitor that diffract to 1.7A resolution were obtained using hanging drop method by vapor diffusion at 18 degrees C. The refined structure shows the conservative beta-trefoil fold features of the Kunitz inhibitors. In BbCI, one of the two characteristic S-S bonds is replaced by the water-mediated interaction between Tyr125 and Gly132. In this work we explore the structural differences between Kunitz-type inhibitors and analyze the essential interactions that maintain the protein structural stability preserving its biological function.
- Departamento de Bioquímica, Universidade Federal de São Paulo-Escola Paulista de Medicina, Rua Três de Maio, 100, 04044-020 São Paulo, SP, Brazil.
Organizational Affiliation: