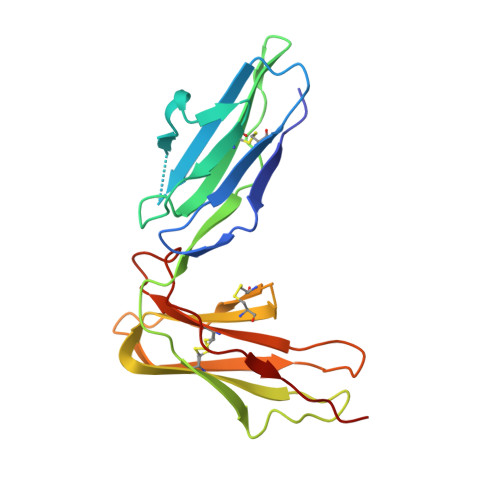
Crystal structure of LIR-2 (ILT4) at 1.8 A: differences from LIR-1 (ILT2) in regions implicated in the binding of the Human Cytomegalovirus class I MHC homolog UL18.
Willcox, B.E., Thomas, L.M., Chapman, T.L., Heikema, A.P., West, A.P., Bjorkman, P.J.(2002) BMC Struct Biol 2: 6-6
- PubMed: 12390682 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1186/1472-6807-2-6
- Primary Citation Related Structures:
2GW5 - PubMed Abstract:
Leukocyte Immunoglobulin-like Receptor-1 (LIR-1) and LIR-2 (also known as ILT2 and ILT4 respectively) are highly related cell surface receptors that bind a broad range of class I MHC molecules with low (microM) affinities. Expressed on monocytic cells and macrophages, both molecules transmit inhibitory signals after binding ligands. In addition to binding host class I MHC, the LIR-1 molecule, which is also expressed on lymphoid tissues, binds with a high (nM) affinity to UL18, a class I MHC homolog encoded by Human Cytomegalovirus (HCMV). In comparison, LIR-2 binds UL18 only weakly (microM KD). To understand how HCMV preferentially targets the more broadly expressed LIR-1 molecule, we determined the crystal structure of a ligand-binding fragment of LIR-2, and compared this to the existing high-resolution crystal structure of LIR-1. Recombinant LIR-2 (domains 1 and 2) was produced in E. coli and crystallized using streak seeding to optimize the crystal morphology. A data set complete to 1.8 A was collected at 100 K from a single crystal in the P4(1)2(1)2 spacegroup. The structure was solved by molecular replacement, using a search model based on the LIR-1 structure. The overall structure of LIR-2 D1D2 resembles both LIR-1, and Killer Inhibitory Receptors, in that the A strand in each domain forms hydrogen bonds to both beta sheets, and there is a sharp angle between the two immunoglobulin-like domains. However, differences from LIR-1 are observed in each domain, with two key changes apparent in the ligand-binding domain, D1. The region corresponding to the residue 44-57 helix of LIR-1 adopts a topology distinct from that of both LIR-1 and the KIR structures, involving a shortened 310 helix. Secondly, the predicted UL18 binding region of LIR-1 is altered substantially in LIR-2: the 76-84 loop mainchain is displaced 11 A with respect to LIR-1, and Tyrosine 38 adopts an alternative rotamer conformation. In summary, the structure of LIR-2 has revealed significant differences to LIR-1, including ones that may help to explain the >1000-fold lower affinity of LIR-2 for UL18.
- Division of Biology 156-29, California Institute of Technology, Pasadena, California 91125, USA. b.willcox@bham.ac.uk
Organizational Affiliation: