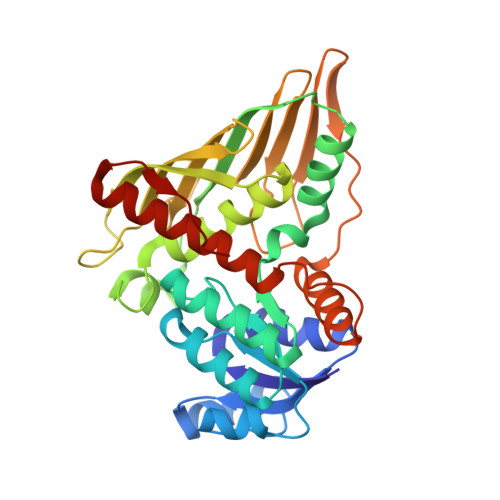
Crystal Structure of NADP(H)-Dependent 1,5-Anhydro-d-fructose Reductase from Sinorhizobium morelense at 2.2 A Resolution: Construction of a NADH-Accepting Mutant and Its Application in Rare Sugar Synthesis
Dambe, T.R., Kuehn, A.M., Brossette, T., Giffhorn, F., Scheidig, A.J.(2006) Biochemistry 45: 10030-10042
- PubMed: 16906761 Search on PubMed
- DOI: https://doi.org/10.1021/bi052589q
- Primary Citation Related Structures:
2GLX - PubMed Abstract:
Recombinant 1,5-anhydro-d-fructose reductase (AFR) from Sinorhizobium morelense S-30.7.5 was crystallized in complex with the cofactor NADP(H) and its structure determined to 2.2 A resolution using selenomethionine SAD (refined R(work) and R(free) factors of 18.9 and 25.0%, respectively). As predicted from the sequence and shown by the structure, AFR can be assigned to the GFO/IDH/MocA protein family. AFR consists of two domains. The N-terminal domain displays a Rossmann fold and contains the cofactor binding site. The intact crystals contain the oxidized cofactor NADP(+), whose attachment to the cofactor binding site is similar to that of NADP(+) in glucose-fructose oxidoreductase (GFOR) from Zymomonas mobilis. Due to variations in length and sequence within loop regions L3 and L5, respectively, the adenine moiety of NADP(+) adopts a different orientation in AFR caused by residue Arg38 forming hydrogen bonds with the 2'-phosphate moiety of NADP(+) and cation-pi stacking interactions with the adenine ring. Amino acid replacements in AFR (S10G, A13G, and S33D) showed that Ala13 is crucial for the discrimination between NADPH and NADH and yielded the A13G variant with dual cosubstrate specificity. The C-terminal domain contains the putative substrate binding site that was occupied by an acetate ion. As determined by analogy to GFOR and by site-directed mutagenesis of K94G, D176A, and H180A, residues Lys94, Asp176, and His180 are most likely involved in substrate binding and catalysis, as substitution of any of these residues resulted in a significant decrease in k(cat) for 1,5-AF. In this context, His180 might serve as a general acid-base catalyst by polarizing the carbonyl function of 1,5-AF to enable the transfer of the hydride from NADPH to the substrate. Here we present the first structure of an AFR enzyme catalyzing the stereoselective reduction of 1,5-AF to 1,5-anhydro-d-mannitol, the final step of a modified anhydrofructose pathway in S. morelense S-30.7.5. We also emphasize the importance of the A13G variant in biocatalysis for the synthesis of 1,5-AM and related derivatives.
- Abteilung Strukturbiologie, Fachbereich Biophysik, Universitätsklinikum des Saarlandes, D-66424 Homburg/Saar, Germany.
Organizational Affiliation: