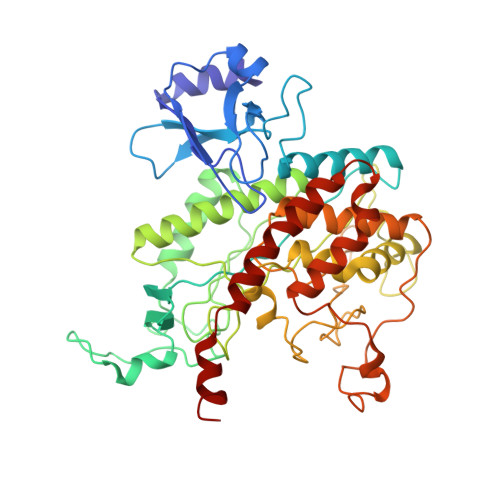
Refined atomic model of glutamine synthetase at 3.5 A resolution.
Yamashita, M.M., Almassy, R.J., Janson, C.A., Cascio, D., Eisenberg, D.(1989) J Biological Chem 264: 17681-17690
- PubMed: 2572586 Search on PubMed
- DOI: https://doi.org/10.2210/pdb2gls/pdb
- Primary Citation Related Structures:
2GLS - PubMed Abstract:
An atomic model of 43,692 non-hydrogen atoms has been determined for the 12-subunit enzyme glutamine synthetase from Salmonella typhimurium, by methods of x-ray diffraction including restrained least-squares atomic refinement against 65,223 unique reflections. At 3.5 A resolution the crystallographic R-factor (on 2 sigma data) is 25.8%. As reported earlier for the unrefined structure, the 12 subunits are arranged in two layers of six; at the interface of pairs of subunits within each layer, cylindrical active sites are formed by six anti-parallel beta strands contributed by one subunit and two strands by the neighboring subunit. This interpretation of the electron density map has now been supported by comparison with glutamine synthetase from Escherichia coli by the Fourier difference method. Each active site cylinder holds two Mn2+ ions, with each ion having as ligands three protein side chains and two water molecules (one water shared by both metals), as well as a histidyl side chain just beyond liganding distance. The protein ligands to Mn2+ 469 are Glu-131, Glu-212, and Glu-220; those to Mn2+ 470 are Glu-129, His-269, and Glu-357. The two layers of subunits are held together largely by the apolar COOH terminus, a helical thong, which inserts into a hydrophobic pocket formed by two neighboring subunits on the opposite ring. Also between layers, there is a hydrogen-bonded beta sheet interaction, as there is between subunits within a ring, but hydrophobic interactions account for most of the intersubunit stability. The central loop, which extends into the central aqueous channel, is subject to attack by at least five enzymes and is discussed as an enzyme "passive site."
- Molecular Biology Institute, University of California, Los Angeles 90024.
Organizational Affiliation: